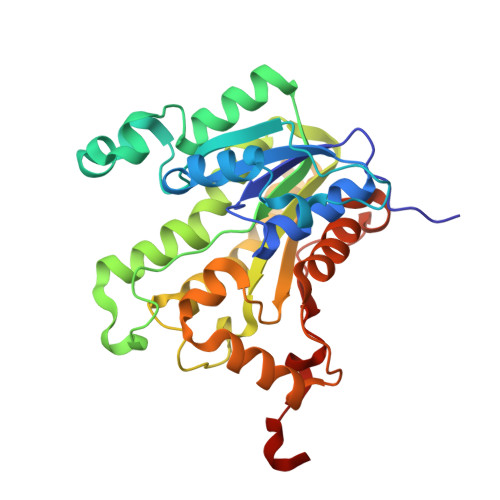
Type II Fatty Acid Synthesis Is Essential for the Replication of Chlamydia trachomatis.
Yao, J., Abdelrahman, Y.M., Robertson, R.M., Cox, J.V., Belland, R.J., White, S.W., Rock, C.O.(2014) J Biological Chem 289: 22365-22376
- PubMed: 24958721 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.584185
- Primary Citation Related Structures:
4Q9N - PubMed Abstract:
The major phospholipid classes of the obligate intracellular bacterial parasite Chlamydia trachomatis are the same as its eukaryotic host except that they also contain chlamydia-made branched-chain fatty acids in the 2-position. Genomic analysis predicts that C. trachomatis is capable of type II fatty acid synthesis (FASII). AFN-1252 was deployed as a chemical tool to specifically inhibit the enoyl-acyl carrier protein reductase (FabI) of C. trachomatis to determine whether chlamydial FASII is essential for replication within the host. The C. trachomatis FabI (CtFabI) is a homotetramer and exhibited typical FabI kinetics, and its expression complemented an Escherichia coli fabI(Ts) strain. AFN-1252 inhibited CtFabI by binding to the FabI·NADH complex with an IC50 of 0.9 μM at saturating substrate concentration. The x-ray crystal structure of the CtFabI·NADH·AFN-1252 ternary complex revealed the specific interactions between the drug, protein, and cofactor within the substrate binding site. AFN-1252 treatment of C. trachomatis-infected HeLa cells at any point in the infectious cycle caused a decrease in infectious titers that correlated with a decrease in branched-chain fatty acid biosynthesis. AFN-1252 treatment at the time of infection prevented the first cell division of C. trachomatis, although the cell morphology suggested differentiation into a metabolically active reticulate body. These results demonstrate that FASII activity is essential for C. trachomatis proliferation within its eukaryotic host and validate CtFabI as a therapeutic target against C. trachomatis.
- From the Departments of Infectious Diseases and.
Organizational Affiliation: