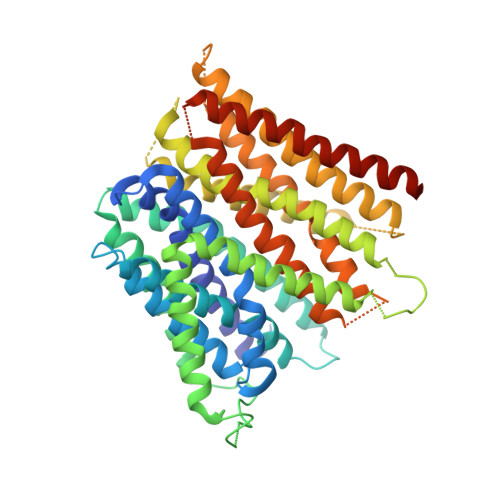
Crystal structure of the E. coli peptide transporter YbgH.
Zhao, Y., Mao, G., Liu, M., Zhang, L., Wang, X., Zhang, X.C.(2014) Structure 22: 1152-1160
- PubMed: 25066136 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2014.06.008
- Primary Citation Related Structures:
4Q65 - PubMed Abstract:
E. coli YbgH belongs to the family of proton-dependent oligopeptide transporters (POTs), a subfamily of the major facilitator superfamily (MFS) of secondary active transporters. Like other MFS transporters, POT proteins switch between two major conformations during substrate transport. Apart from possessing a canonical 12-helix, two-domain transmembrane (TM) core, prokaryotic POT proteins usually have two TM helices inserted between the two domains. Here we determined the crystal structure of YbgH in its inward-facing conformation. Our structure-based functional studies investigated the roles of both the POT signature motif 2 and the inserted interdomain TM helix pair in the stabilization and regulation of the major conformational change in MFS/POT transporters. Furthermore, of all the proton-titratable amino acid residues, Glu21 is the only conserved one (among POTs) located in the central cavity and is critical for in vivo transport. Together, our results support the notion that MFS symporters utilize a transport mechanism based on substrate-protonation coupling.
- School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, China; National Laboratory of Macromolecules, National Center of Protein Science Beijing, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing 100101, China.
Organizational Affiliation: