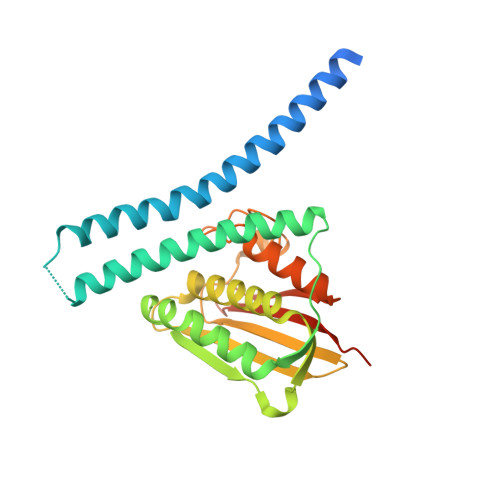
Cell fate regulation governed by a repurposed bacterial histidine kinase.
Childers, W.S., Xu, Q., Mann, T.H., Mathews, I.I., Blair, J.A., Deacon, A.M., Shapiro, L.(2014) PLoS Biol 12: e1001979-e1001979
- PubMed: 25349992 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1001979
- Primary Citation Related Structures:
4Q20 - PubMed Abstract:
One of the simplest organisms to divide asymmetrically is the bacterium Caulobacter crescentus. The DivL pseudo-histidine kinase, positioned at one cell pole, regulates cell-fate by controlling the activation of the global transcription factor CtrA via an interaction with the response regulator (RR) DivK. DivL uniquely contains a tyrosine at the histidine phosphorylation site, and can achieve these regulatory functions in vivo without kinase activity. Determination of the DivL crystal structure and biochemical analysis of wild-type and site-specific DivL mutants revealed that the DivL PAS domains regulate binding specificity for DivK∼P over DivK, which is modulated by an allosteric intramolecular interaction between adjacent domains. We discovered that DivL's catalytic domains have been repurposed as a phosphospecific RR input sensor, thereby reversing the flow of information observed in conventional histidine kinase (HK)-RR systems and coupling a complex network of signaling proteins for cell-fate regulation.
- Department of Developmental Biology, Stanford University School of Medicine, Stanford, California, United States of America.
Organizational Affiliation: