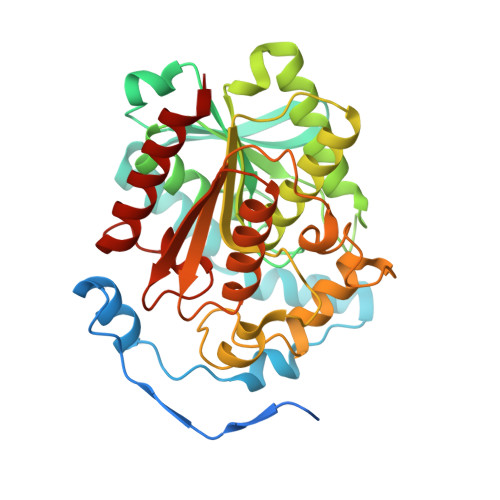
Structural basis for dimerization and catalysis of a novel esterase from the GTSAG motif subfamily of the bacterial hormone-sensitive lipase family
Li, P.Y., Ji, P., Li, C.Y., Zhang, Y., Wang, G.L., Zhang, X.Y., Xie, B.B., Qin, Q.L., Chen, X.L., Zhou, B.C., Zhang, Y.Z.(2014) J Biological Chem 289: 19031-19041
- PubMed: 24867954
- DOI: https://doi.org/10.1074/jbc.M114.574913
- Primary Citation of Related Structures:
4Q05 - PubMed Abstract:
Hormone-sensitive lipases (HSLs) are widely distributed in microorganisms, plants, and animals. Microbial HSLs are classified into two subfamilies, an unnamed new subfamily and the GDSAG motif subfamily. Due to the lack of structural information, the detailed catalytic mechanism of the new subfamily is not yet clarified. Based on sequence analysis, we propose to name the new subfamily as the GTSAG motif subfamily. We identified a novel HSL esterase E25, a member of the GTSAG motif subfamily, by functional metagenomic screening, and resolved its structure at 2.05 Å. E25 is mesophilic (optimum temperature at 50 °C), salt-tolerant, slightly alkaline (optimum pH at 8.5) for its activity, and capable of hydrolyzing short chain monoesters (C2-C10). E25 tends to form dimers both in the crystal and in solution. An E25 monomer contains an N-terminal CAP domain, and a classical α/β hydrolase-fold domain. Residues Ser(186), Asp(282), and His(312) comprise the catalytic triad. Structural and mutational analyses indicated that E25 adopts a dimerization pattern distinct from other HSLs. E25 dimer is mainly stabilized by an N-terminal loop intersection from the CAP domains and hydrogen bonds and salt bridges involving seven highly conserved hydrophilic residues from the catalytic domains. Further analysis indicated that E25 also has some catalytic profiles different from other HSLs. Dimerization is essential for E25 to exert its catalytic activity by keeping the accurate orientation of the catalytic Asp(282) within the catalytic triad. Our results reveal the structural basis for dimerization and catalysis of an esterase from the GTSAG motif subfamily of the HSL family.
- From the State Key Laboratory of Microbial Technology, Marine Biotechnology Research Center.
Organizational Affiliation: