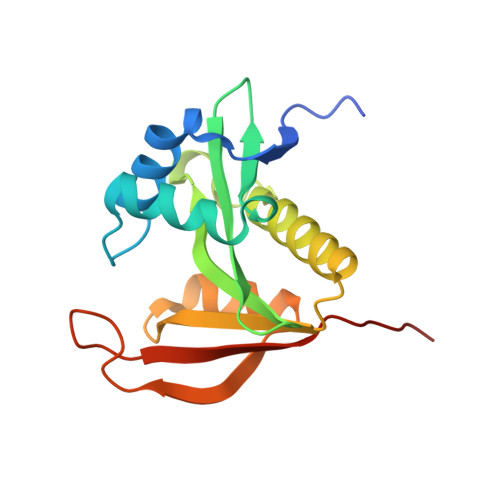
Structure of Thermoplasma volcanium Ard1 belongs to N-acetyltransferase family member suggesting multiple ligand binding modes with acetyl coenzyme A and coenzyme A.
Ma, C., Pathak, C., Jang, S., Lee, S.J., Nam, M., Kim, S.J., Im, H., Lee, B.J.(2014) Biochim Biophys Acta 1844: 1790-1797
- PubMed: 25062911
- DOI: https://doi.org/10.1016/j.bbapap.2014.07.011
- Primary Citation of Related Structures:
4PV6 - PubMed Abstract:
Acetylation and deacetylation reactions result in biologically important modifications that are involved in normal cell function and cancer development. These reactions, carried out by protein acetyltransferase enzymes, act by transferring an acetyl group from acetyl-coenzymeA (Ac-CoA) to various substrate proteins. Such protein acetylation remains poorly understood in Archaea, and has been only partially described. Information processing in Archaea has been reported to be similar to that in eukaryotes and distinct from the equivalent bacterial processes. The human N-acetyltransferase Ard1 (hArd1) is one of the acetyltransferases that has been found to be overexpressed in various cancer cells and tissues, and knockout of the hArd1 gene significantly reduces growth rate of the cancer cell lines. In the present study, we determined the crystal structure of Thermoplasma volcanium Ard1 (Tv Ard1), which shows both ligand-free and multiple ligand-bound forms, i.e.,Ac-CoA- and coenzyme A (CoA)-bound forms. The difference between ligand-free and ligand-bound chains in the crystal structure was used to search for the interacting residues. The re-orientation and position of the loop between β4 and α3 including the phosphate-binding loop (P-loop) were observed, which are important for the ligand interaction. In addition, a biochemical assay to determine the N-acetyltransferase activity of Tv Ard1 was performed using the T.volcanium substrate protein Alba (Tv Alba). Taken together, the findings of this study elucidate ligand-free form of Tv Ard1 for the first time and suggest multiple modes of binding with Ac-CoA and CoA.
- Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Gwanak-gu, Seoul 151-742, Republic of Korea.
Organizational Affiliation: