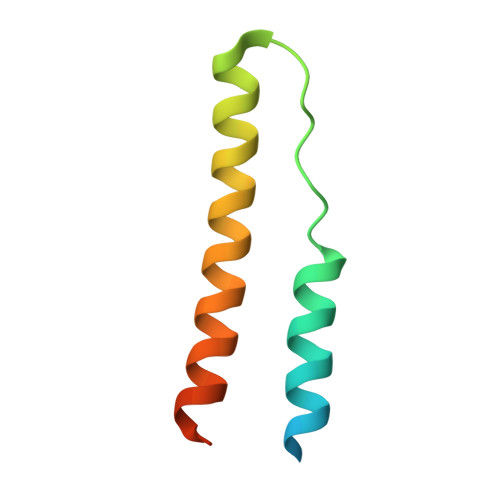
RNA-directed remodeling of the HIV-1 protein Rev orchestrates assembly of the Rev-Rev response element complex.
Jayaraman, B., Crosby, D.C., Homer, C., Ribeiro, I., Mavor, D., Frankel, A.D.(2014) Elife 4: e04120-e04120
- PubMed: 25486594 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.04120
- Primary Citation Related Structures:
4PMI - PubMed Abstract:
The HIV-1 protein Rev controls a critical step in viral replication by mediating the nuclear export of unspliced and singly-spliced viral mRNAs. Multiple Rev subunits assemble on the Rev Response Element (RRE), a structured region present in these RNAs, and direct their export through the Crm1 pathway. Rev-RRE assembly occurs via several Rev oligomerization and RNA-binding steps, but how these steps are coordinated to form an export-competent complex is unclear. Here, we report the first crystal structure of a Rev dimer-RRE complex, revealing a dramatic rearrangement of the Rev-dimer upon RRE binding through re-packing of its hydrophobic protein-protein interface. Rev-RNA recognition relies on sequence-specific contacts at the well-characterized IIB site and local RNA architecture at the second site. The structure supports a model in which the RRE utilizes the inherent plasticity of Rev subunit interfaces to guide the formation of a functional complex.
- Department of Biochemistry and Biophysics, University of California, San Francisco, San Francisco, United States.
Organizational Affiliation: