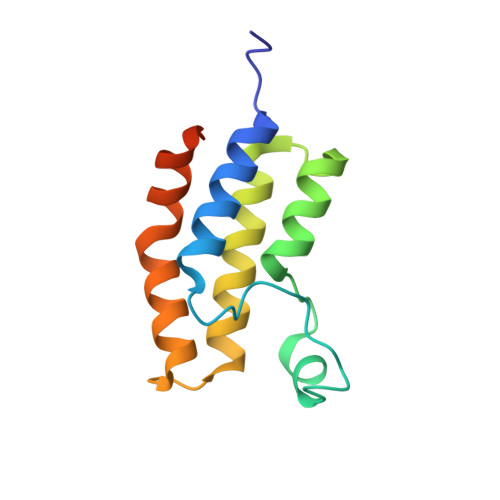
Bromodomain Proteins Contribute to Maintenance of Bloodstream Form Stage Identity in the African Trypanosome.
Schulz, D., Mugnier, M.R., Paulsen, E.M., Kim, H.S., Chung, C.W., Tough, D.F., Rioja, I., Prinjha, R.K., Papavasiliou, F.N., Debler, E.W.(2015) PLoS Biol 13: e1002316-e1002316
- PubMed: 26646171 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1002316
- Primary Citation Related Structures:
4PKL - PubMed Abstract:
Trypanosoma brucei, the causative agent of African sleeping sickness, is transmitted to its mammalian host by the tsetse. In the fly, the parasite's surface is covered with invariant procyclin, while in the mammal it resides extracellularly in its bloodstream form (BF) and is densely covered with highly immunogenic Variant Surface Glycoprotein (VSG). In the BF, the parasite varies this highly immunogenic surface VSG using a repertoire of ~2500 distinct VSG genes. Recent reports in mammalian systems point to a role for histone acetyl-lysine recognizing bromodomain proteins in the maintenance of stem cell fate, leading us to hypothesize that bromodomain proteins may maintain the BF cell fate in trypanosomes. Using small-molecule inhibitors and genetic mutants for individual bromodomain proteins, we performed RNA-seq experiments that revealed changes in the transcriptome similar to those seen in cells differentiating from the BF to the insect stage. This was recapitulated at the protein level by the appearance of insect-stage proteins on the cell surface. Furthermore, bromodomain inhibition disrupts two major BF-specific immune evasion mechanisms that trypanosomes harness to evade mammalian host antibody responses. First, monoallelic expression of the antigenically varied VSG is disrupted. Second, rapid internalization of antibodies bound to VSG on the surface of the trypanosome is blocked. Thus, our studies reveal a role for trypanosome bromodomain proteins in maintaining bloodstream stage identity and immune evasion. Importantly, bromodomain inhibition leads to a decrease in virulence in a mouse model of infection, establishing these proteins as potential therapeutic drug targets for trypanosomiasis. Our 1.25Å resolution crystal structure of a trypanosome bromodomain in complex with I-BET151 reveals a novel binding mode of the inhibitor, which serves as a promising starting point for rational drug design.
- Laboratory of Lymphocyte Biology, The Rockefeller University, New York, New York, United States of America.
Organizational Affiliation: