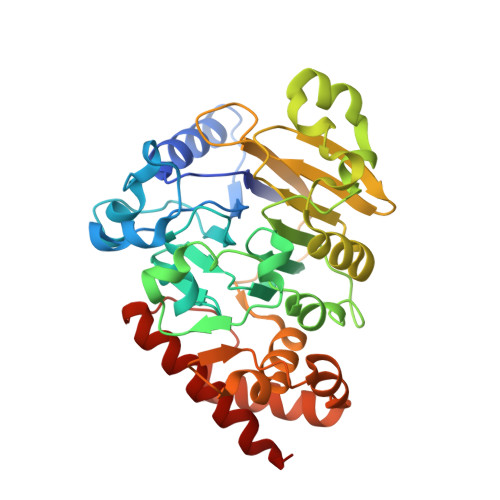
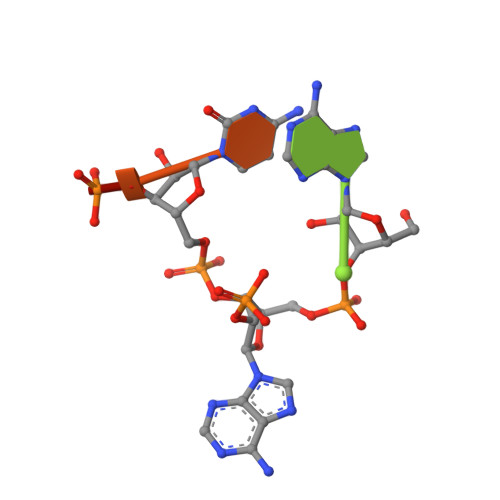
Structural basis of lariat RNA recognition by the intron debranching enzyme Dbr1.
Montemayor, E.J., Katolik, A., Clark, N.E., Taylor, A.B., Schuermann, J.P., Combs, D.J., Johnsson, R., Holloway, S.P., Stevens, S.W., Damha, M.J., Hart, P.J.(2014) Nucleic Acids Res 42: 10845-10855
- PubMed: 25123664
- DOI: https://doi.org/10.1093/nar/gku725
- Primary Citation of Related Structures:
4PEF, 4PEG, 4PEH, 4PEI - PubMed Abstract:
The enzymatic processing of cellular RNA molecules requires selective recognition of unique chemical and topological features. The unusual 2',5'-phosphodiester linkages in RNA lariats produced by the spliceosome must be hydrolyzed by the intron debranching enzyme (Dbr1) before they can be metabolized or processed into essential cellular factors, such as snoRNA and miRNA. Dbr1 is also involved in the propagation of retrotransposons and retroviruses, although the precise role played by the enzyme in these processes is poorly understood. Here, we report the first structures of Dbr1 alone and in complex with several synthetic RNA compounds that mimic the branchpoint in lariat RNA. The structures, together with functional data on Dbr1 variants, reveal the molecular basis for 2',5'-phosphodiester recognition and explain why the enzyme lacks activity toward 3',5'-phosphodiester linkages. The findings illuminate structure/function relationships in a unique enzyme that is central to eukaryotic RNA metabolism and set the stage for the rational design of inhibitors that may represent novel therapeutic agents to treat retroviral infections and neurodegenerative disease.
- Department of Biochemistry, The University of Texas Health Science Center, San Antonio, TX 78229, USA X-ray Crystallography Core Laboratory, The University of Texas Health Science Center, San Antonio, TX 78229, USA.
Organizational Affiliation: