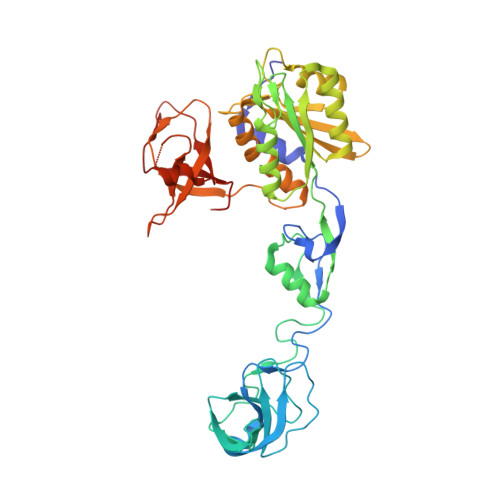
Modulation of gephyrin-glycine receptor affinity by multivalency.
Maric, H.M., Kasaragod, V.B., Schindelin, H.(2014) ACS Chem Biol 9: 2554-2562
- PubMed: 25137389 Search on PubMed
- DOI: https://doi.org/10.1021/cb500303a
- Primary Citation Related Structures:
4PD0, 4PD1 - PubMed Abstract:
Gephyrin is a major determinant for the accumulation and anchoring of glycine receptors (GlyRs) and the majority of γ-aminobutyric acid type A receptors (GABAARs) at postsynaptic sites. Here we explored the interaction of gephyrin with a dimeric form of a GlyR β-subunit receptor-derived peptide. A 2 Å crystal structure of the C-terminal domain of gephyrin (GephE) in complex with a 15-residue peptide derived from the GlyR β-subunit defined the core binding site, which we targeted with the dimeric peptide. Biophysical analyses via differential scanning calorimetry (DSC), thermofluor, and isothermal titration calorimetry (ITC) demonstrated that this dimeric ligand is capable of binding simultaneously to two receptor binding sites and that this multivalency results in a 25-fold enhanced affinity. Our study therefore suggests that the oligomeric state of gephyrin and the number of gephyrin-binding subunits in the pentameric GABAARs and GlyRs together control postsynaptic receptor clustering.
- Institute of Structural Biology, Rudolf Virchow Center for Experimental Biomedicine, University of Würzburg , Josef-Schneider-Str. 2, 97080 Würzburg, Germany.
Organizational Affiliation: