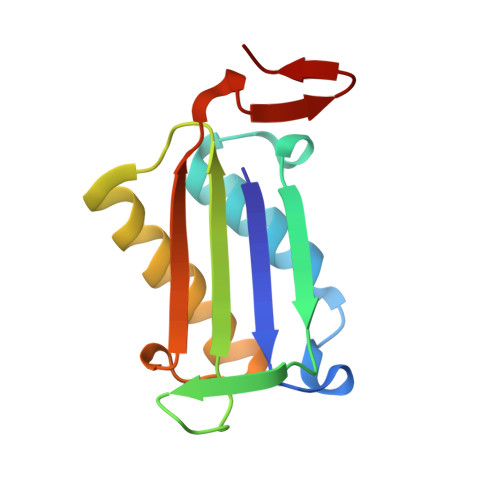
(6-isothiocyanatohexyl)benzene inhibitor complexed with Macrophage Migration Inhibitory Factor
Spencer, E.S., Dale, E.J., Gommans, A.L., Vo, C.T., Rutledge, M.T., Nakatani, Y., Gamble, A.B., Smith, R.A.J., Wilbanks, S.M., Hampton, M.B., Tyndall, J.D.A.To be published.