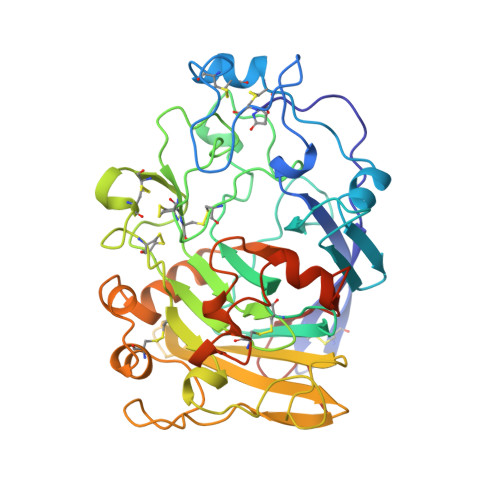
Structure of the endoglucanase I from Fusarium oxysporum: native, cellobiose, and 3,4-epoxybutyl beta-D-cellobioside-inhibited forms, at 2.3 A resolution.
Sulzenbacher, G., Schulein, M., Davies, G.J.(1997) Biochemistry 36: 5902-5911
- PubMed: 9153432
- DOI: https://doi.org/10.1021/bi962963+
- Primary Citation of Related Structures:
2OVW, 3OVW, 4OVW - PubMed Abstract:
The mechanisms involved in the enzymatic degradation of cellulose are of great ecological and commercial importance. The breakdown of cellulose by fungal species is performed by a consortium of free enzymes, known as cellobiohydrolases and endoglucanases, which are found in many of the 57 glycosyl hydrolase families. The structure of the endoglucanase I (EG I), found in glycosyl hydrolase family 7, from the thermophilic fungus Fusarium oxysporum has been solved at 2.3 A resolution. In addition to the native enzyme, structures have also been determined with both the affinity label, 3,4-epoxybutyl beta-D-cellobioside, and the reaction product cellobiose. The affinity label is covalently bound, as expected, to the catalytic nucleophile, Glu197, with clear evidence for binding of both the R and S stereoisomers. Cellobiose is found bound to the -2 and -1 subsites of the enzyme. In marked contrast to the structure of EG I with a nonhydrolyzable thiosaccharide analog, which spanned the -2, -1, and +1 subsites and which had a skew-boat conformation for the -1 subsite sugar [Sulzenbacher, G., et al. (1996) Biochemistry 35, 15280-15287], the cellobiose complex shows no pyranoside ring distortion in the -1 subsite, implying that strain is induced primarily by the additional +1 subsite interactions and that the product is found, as expected, in its unstrained conformation.
- Department of Chemistry, University of York, Heslington, U.K.
Organizational Affiliation: