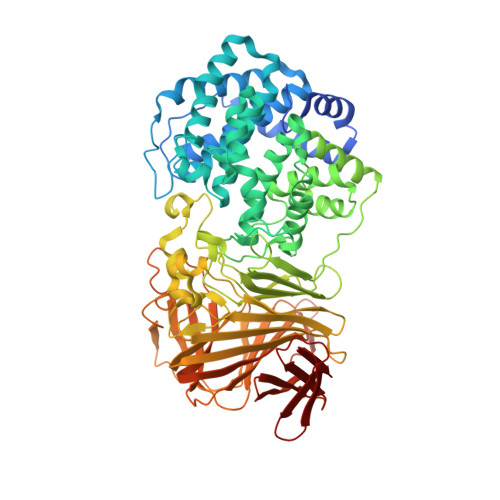
Structure of a PL17 Family Alginate Lyase Demonstrates Functional Similarities among Exotype Depolymerases.
Park, D., Jagtap, S., Nair, S.K.(2014) J Biological Chem 289: 8645-8655
- PubMed: 24478312
- DOI: https://doi.org/10.1074/jbc.M113.531111
- Primary Citation Related Structures:
4NEI, 4OJZ, 4OK2, 4OK4 - PubMed Abstract:
Brown macroalgae represent an ideal source for complex polysaccharides that can be utilized as precursors for cellulosic biofuels. The lack of recalcitrant lignin components in macroalgae polysaccharide reserves provides a facile route for depolymerization of constituent polysaccharides into simple monosaccharides. The most abundant sugars in macroalgae are alginate, mannitol, and glucan, and although several classes of enzymes that can catabolize the latter two have been characterized, studies of alginate-depolymerizing enzymes have lagged. Here, we present several crystal structures of Alg17c from marine bacterium Saccharophagus degradans along with structure-function characterization of active site residues that are suggested to be involved in the exolytic mechanism of alginate depolymerization. This represents the first structural and biochemical characterization of a family 17 polysaccharide lyase enzyme. Despite the lack of appreciable sequence conservation, the structure and β-elimination mechanism for glycolytic bond cleavage by Alg17c are similar to those observed for family 15 polysaccharide lyases and other lyases. This work illuminates the evolutionary relationships among enzymes within this unexplored class of polysaccharide lyases and reinforces the notion of a structure-based hierarchy in the classification of these enzymes.
- From the Departments of Biochemistry and.
Organizational Affiliation: