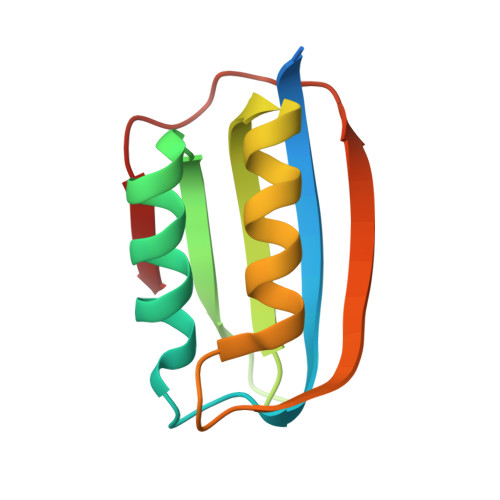
Edge strand engineering prevents native-like aggregation in Sulfolobus solfataricus acylphosphatase.
de Rosa, M., Bemporad, F., Pellegrino, S., Chiti, F., Bolognesi, M., Ricagno, S.(2014) FEBS J 281: 4072-4084
- PubMed: 24893801 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12861
- Primary Citation Related Structures:
4OJ3, 4OJG, 4OJH - PubMed Abstract:
β-proteins are constantly threatened by the risk of aggregation because β-sheets are inherently structured for edge-to-edge interactions. To avoid native-like aggregation, evolution has resulted in a set of strategies that prevent intermolecular β-interactions. Acylphosphatase from Sulfolobus solfataricus (Sso AcP) represents a suitable model for the study of such a process. Under conditions promoting aggregation, Sso AcP acquires a native-like conformational state whereby an unstructured N-terminal segment interacts with the edge β-strand B4 of an adjacent Sso AcP molecule. Because B4 is poorly protected against aggregation, this interaction triggers the aggregation cascade without the need for unfolding. Recently, three single Sso AcP mutants (V84D, Y86E and V84P) were designed to engineer additional protection against aggregation in B4 and were observed to successfully impair native-like aggregation in all three variants at the expense of a lower stability. To understand the structural basis of the reduced aggregation propensity and lower stability, the crystal structures of the Sso AcP variants were determined in the present study. Structural analysis reveals that the V84D and Y86E mutations exert protection by the insertion of an edge negative charge. A conformationally less regular B4 underlies protection against aggregation in the V84P mutant. The thermodynamic basis of instability is discussed. Moreover, kinetic experiments indicate that aggregation of the three mutants is not native-like and is independent of the interaction between B4 and the unstructured N-terminal segment. The reported data rationalize previous evidence regarding Sso AcP native-like aggregation and provide a basis for the design of aggregation-free proteins. The atomic coordinates and related experimental data for the Sso AcP mutants V84P, V84D, ΔN11 Y86E have been deposited in the Protein Data Bank under accession numbers 4OJ3, 4OJG and 4OJH, respectively. • Sso AcP and Sso AcP bind by fluorescence technology (View interaction).
- Dipartimento di Bioscienze, Università di Milano, Italy.
Organizational Affiliation: