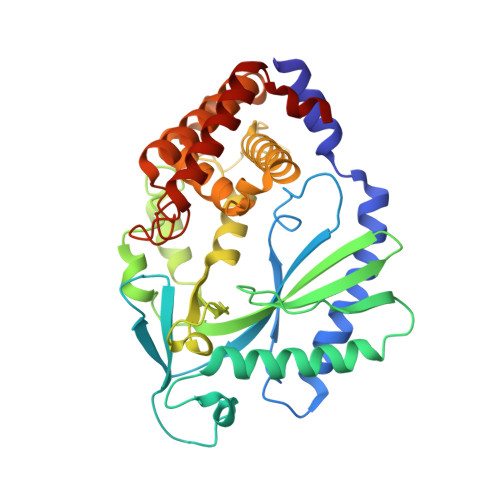
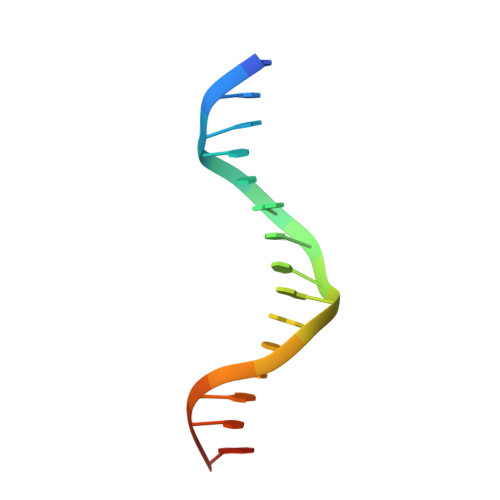
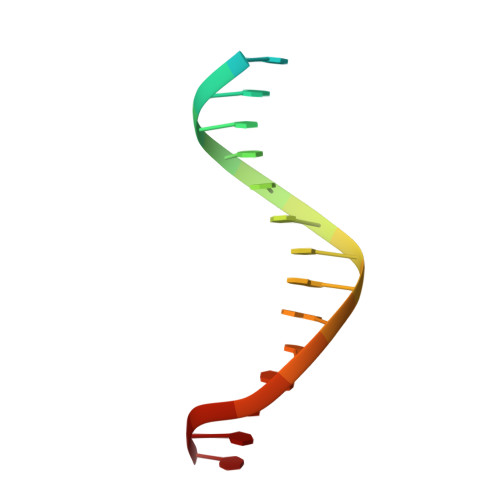
The Cytosolic DNA Sensor cGAS Forms an Oligomeric Complex with DNA and Undergoes Switch-like Conformational Changes in the Activation Loop.
Zhang, X., Wu, J., Du, F., Xu, H., Sun, L., Chen, Z., Brautigam, C.A., Zhang, X., Chen, Z.J.(2014) Cell Rep 6: 421-430
- PubMed: 24462292 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2014.01.003
- Primary Citation Related Structures:
4O67, 4O68, 4O69, 4O6A - PubMed Abstract:
The presence of DNA in the cytoplasm is a danger signal that triggers immune and inflammatory responses. Cytosolic DNA binds to and activates cyclic GMP-AMP (cGAMP) synthase (cGAS), which produces the second messenger cGAMP. cGAMP binds to the adaptor protein STING and activates a signaling cascade that leads to the production of type I interferons and other cytokines. Here, we report the crystal structures of human cGAS in its apo form, representing its autoinhibited conformation as well as in its cGAMP- and sulfate-bound forms. These structures reveal switch-like conformational changes of an activation loop that result in the rearrangement of the catalytic site. The structure of DNA-bound cGAS reveals a complex composed of dimeric cGAS bound to two molecules of DNA. Functional analyses of cGAS mutants demonstrate that both the protein-protein interface and the two DNA binding surfaces are critical for cGAS activation. These results provide insights into the mechanism of DNA sensing by cGAS.
- Department of Molecular Biology, University of Texas Southwestern Medical Center, Dallas, TX 75390-9148, USA. Electronic address: xu.zhang@utsouthwestern.edu.
Organizational Affiliation: