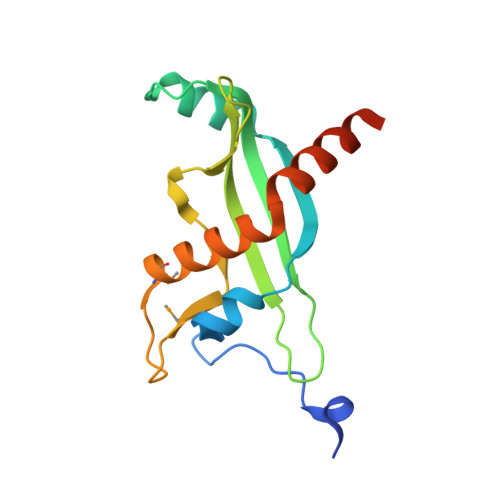
RsaM: a transcriptional regulator of Burkholderia spp. with novel fold.
Michalska, K., Chhor, G., Clancy, S., Jedrzejczak, R., Babnigg, G., Winans, S.C., Joachimiak, A.(2014) FEBS J 281: 4293-4306
- PubMed: 24916958 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.12868
- Primary Citation Related Structures:
4O2H - PubMed Abstract:
Burkholderia cepacia complex is a set of closely related bacterial species that are notorious pathogens of cystic fibrosis patients, responsible for life-threatening lung infections. Expression of several virulence factors of Burkholderia cepacia complex is controlled by a mechanism known as quorum sensing (QS). QS is a means of bacterial communication used to coordinate gene expression in a cell-density-dependent manner. The system involves the production of diffusible signaling molecules (N-acyl-l-homoserine lactones, AHLs), that bind to cognate transcriptional regulators and influence their ability to regulate gene expression. One such system that is highly conserved in Burkholderia cepacia complex consists of CepI and CepR. CepI is AHL synthase, whereas CepR is an AHL-dependent transcription factor. In most members of the Burkholderia cepacia complex group, the cepI and cepR genes are divergently transcribed and separated by additional genes. One of them, bcam1869, encodes the BcRsaM protein, which was recently postulated to modulate the abundance or activity of CepI or CepR. Here, we show the crystal structure of BcRsaM from B. cenocepacia J2315. It is a single-domain protein with unique topology and presents a novel fold. The protein is a dimer in the crystal and in solution. This regulator has no known DNA-binding motifs and direct binding of BcRsaM to the cepI promoter could not be detected in in vitro assays. Therefore, we propose that the modulatory action of RsaM might result from interactions with other components of the QS machinery rather than from direct association with the DNA promoter. The atomic coordinates and structure factors have been deposited in the Protein Data Bank under entry 4O2H. BcRsaM and BcRsaM bind by x-ray crystallography (View interaction) BcRsaM and BcRsaM bind by molecular sieving (View interaction).
- Midwest Center for Structural Genomics, Argonne National Laboratory, IL, USA; Structural Biology Center, Biosciences Division, Argonne National Laboratory, IL, USA.
Organizational Affiliation: