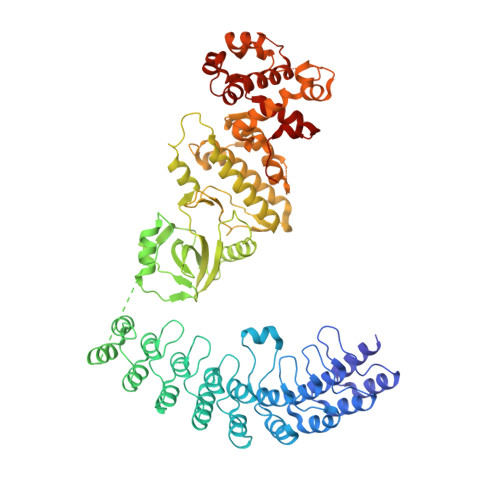
Dimeric structure of pseudokinase RNase L bound to 2-5A reveals a basis for interferon-induced antiviral activity.
Huang, H., Zeqiraj, E., Dong, B., Jha, B.K., Duffy, N.M., Orlicky, S., Thevakumaran, N., Talukdar, M., Pillon, M.C., Ceccarelli, D.F., Wan, L.C., Juang, Y.C., Mao, D.Y., Gaughan, C., Brinton, M.A., Perelygin, A.A., Kourinov, I., Guarne, A., Silverman, R.H., Sicheri, F.(2014) Mol Cell 53: 221-234
- PubMed: 24462203 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2013.12.025
- Primary Citation Related Structures:
4O1O, 4O1P - PubMed Abstract:
RNase L is an ankyrin repeat domain-containing dual endoribonuclease-pseudokinase that is activated by unusual 2,'5'-oligoadenylate (2-5A) second messengers and which impedes viral infections in higher vertebrates. Despite its importance in interferon-regulated antiviral innate immunity, relatively little is known about its precise mechanism of action. Here we present a functional characterization of 2.5 Å and 3.25 Å X-ray crystal and small-angle X-ray scattering structures of RNase L bound to a natural 2-5A activator with and without ADP or the nonhydrolysable ATP mimetic AMP-PNP. These studies reveal how recognition of 2-5A through interactions with the ankyrin repeat domain and the pseudokinase domain, together with nucleotide binding, imposes a rigid intertwined dimer configuration that is essential for RNase catalytic and antiviral functions. The involvement of the pseudokinase domain of RNase L in 2-5A sensing, nucleotide binding, dimerization, and ribonuclease functions highlights the evolutionary adaptability of the eukaryotic protein kinase fold.
- Program in Systems Biology, Lunenfeld-Tanenbaum Research Institute, Mount Sinai Hospital, 600 University Avenue, Toronto, ON M5G 1X5, Canada; Department of Molecular Genetics, University of Toronto, Toronto, ON M5S 1A8, Canada.
Organizational Affiliation: