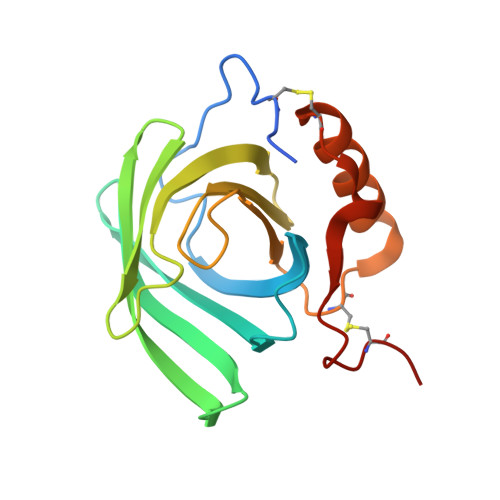
Structure of a heterogeneous, glycosylated, lipid-bound, in vivo-grown protein crystal at atomic resolution from the viviparous cockroach Diploptera punctata.
Banerjee, S., Coussens, N.P., Gallat, F.X., Sathyanarayanan, N., Srikanth, J., Yagi, K.J., Gray, J.S., Tobe, S.S., Stay, B., Chavas, L.M., Ramaswamy, S.(2016) IUCrJ 3: 282-293
- PubMed: 27437115 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252516008903
- Primary Citation Related Structures:
4NYQ, 4NYR, 5EPQ - PubMed Abstract:
Macromolecular crystals for X-ray diffraction studies are typically grown in vitro from pure and homogeneous samples; however, there are examples of protein crystals that have been identified in vivo. Recent developments in micro-crystallography techniques and the advent of X-ray free-electron lasers have allowed the determination of several protein structures from crystals grown in cellulo. Here, an atomic resolution (1.2 Å) crystal structure is reported of heterogeneous milk proteins grown inside a living organism in their functional niche. These in vivo-grown crystals were isolated from the midgut of an embryo within the only known viviparous cockroach, Diploptera punctata. The milk proteins crystallized in space group P1, and a structure was determined by anomalous dispersion from the native S atoms. The data revealed glycosylated proteins that adopt a lipocalin fold, bind lipids and organize to form a tightly packed crystalline lattice. A single crystal is estimated to contain more than three times the energy of an equivalent mass of dairy milk. This unique storage form of nourishment for developing embryos allows access to a constant supply of complete nutrients. Notably, the crystalline cockroach-milk proteins are highly heterogeneous with respect to amino-acid sequence, glycosylation and bound fatty-acid composition. These data present a unique example of protein heterogeneity within a single in vivo-grown crystal of a natural protein in its native environment at atomic resolution.
- Institute of Stem Cell Biology and Regenerative Medicine , Bellary Road, GKVK Campus, Bangalore, Karnataka 560 065, India.
Organizational Affiliation: