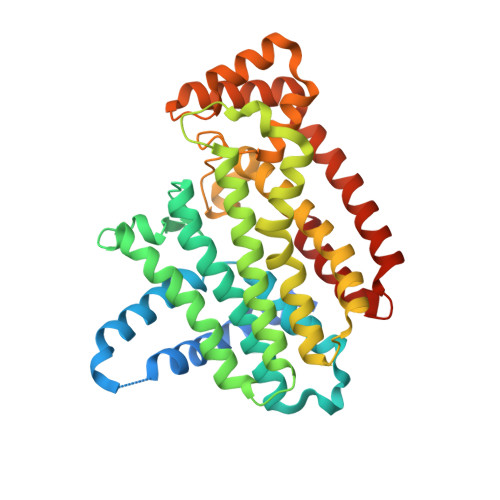
The effects of Lysine 200 and Phenylalanine 239 Farnesyl Pyrophosphate Synthase (FPPS) mutations on the catalytic activity, crystal structure and inhibition by nitrogen containing bisphosphonates
Tsoumpra, M.K., Barnett, B.L., Muniz, J.R.C., Walter, R.L., Ebetino, F.H., von Delft, F., Russell, R.G.G., Oppermann, U., Dunford, J.E.To be published.