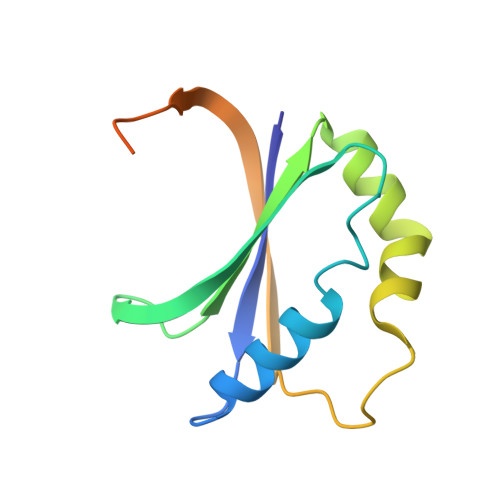
Crystallographic and Spectroscopic Insights into Heme Degradation by Mycobacterium tuberculosis MhuD.
Graves, A.B., Morse, R.P., Chao, A., Iniguez, A., Goulding, C.W., Liptak, M.D.(2014) Inorg Chem 53: 5931-5940
- PubMed: 24901029 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ic500033b
- Primary Citation Related Structures:
4NL5 - PubMed Abstract:
Mycobacterium heme utilization degrader (MhuD) is a heme-degrading protein from Mycobacterium tuberculosis responsible for extracting the essential nutrient iron from host-derived heme. MhuD has been previously shown to produce unique organic products compared to those of canonical heme oxygenases (HOs) as well as those of the IsdG/I heme-degrading enzymes from Staphylococcus aureus. Here, we report the X-ray crystal structure of cyanide-inhibited MhuD (MhuD-heme-CN) as well as detailed (1)H nuclear magnetic resonance (NMR), UV/vis absorption, and magnetic circular dichroism (MCD) spectroscopic characterization of this species. There is no evidence for an ordered network of water molecules on the distal side of the heme substrate in the X-ray crystal structure, as was previously reported for canonical HOs. The degree of heme ruffling in the crystal structure of MhuD is greater than that observed for HO and less than that observed for IsdI. As a consequence, the Fe 3dxz-, 3dyz-, and 3dxy-based MOs are very close in energy, and the room-temperature (1)H NMR spectrum of MhuD-heme-CN is consistent with population of both a (2)Eg electronic state with a (dxy)(2)(dxz,dyz)(3) electron configuration, similar to the ground state of canonical HOs, and a (2)B2g state with a (dxz,dyz)(4)(dxy)(1) electron configuration, similar to the ground state of cyanide-inhibited IsdI. Variable temperature, variable field MCD saturation magnetization data establishes that MhuD-heme-CN has a (2)B2g electronic ground state with a low-lying (2)Eg excited state. Our crystallographic and spectroscopic data suggest that there are both structural and electronic contributions to the α-meso regioselectivity of MhuD-catalyzed heme cleavage. The structural distortion of the heme substrate observed in the X-ray crystal structure of MhuD-heme-CN is likely to favor cleavage at the α- and γ-meso carbons, whereas the spin density distribution may favor selective oxygenation of the α-meso carbon.
- Department of Chemistry, University of Vermont , Burlington, Vermont 05405, United States.
Organizational Affiliation: