Catalytic Mechanism and Mode of Action of the Periplasmic Alginate Epimerase AlgG.
Wolfram, F., Kitova, E.N., Robinson, H., Walvoort, M.T., Codee, J.D., Klassen, J.S., Howell, P.L.(2014) J Biological Chem 289: 6006-6019
- PubMed: 24398681 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.533158
- Primary Citation Related Structures:
4NK6, 4NK8 - PubMed Abstract:
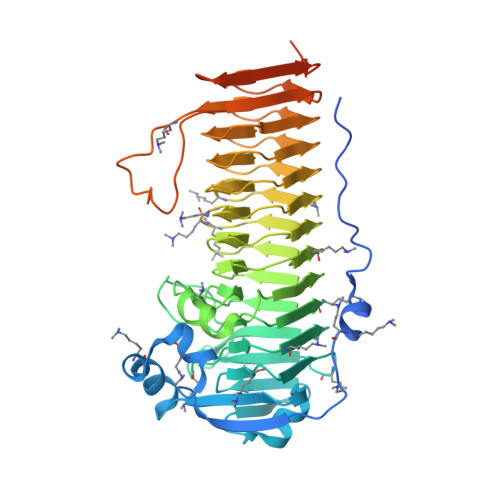
Pseudomonas aeruginosa is an opportunistic pathogen that forms chronic biofilm infections in the lungs of cystic fibrosis patients. A major component of the biofilm during these infections is the exopolysaccharide alginate, which is synthesized at the inner membrane as a homopolymer of 1-4-linked β-D-mannuronate. As the polymer passages through the periplasm, 22-44% of the mannuronate residues are converted to α-L-guluronate by the C5-epimerase AlgG to produce a polymer of alternating β-D-mannuronate and α-L-guluronate blocks and stretches of polymannuronate. To understand the molecular basis of alginate epimerization, the structure of Pseudomonas syringae AlgG has been determined at 2.1-Å resolution, and the protein was functionally characterized. The structure reveals that AlgG is a long right-handed parallel β-helix with an elaborate lid structure. Functional analysis of AlgG mutants suggests that His(319) acts as the catalytic base and that Arg(345) neutralizes the acidic group during the epimerase reaction. Water is the likely catalytic acid. Electrostatic surface potential and residue conservation analyses in conjunction with activity and substrate docking studies suggest that a conserved electropositive groove facilitates polymannuronate binding and contains at least nine substrate binding subsites. These subsites likely align the polymer in the correct register for catalysis to occur. The presence of multiple subsites, the electropositive groove, and the non-random distribution of guluronate in the alginate polymer suggest that AlgG is a processive enzyme. Moreover, comparison of AlgG and the extracellular alginate epimerase AlgE4 of Azotobacter vinelandii provides a structural rationale for the differences in their Ca(2+) dependence.
- From the Program in Molecular Structure and Function, The Hospital for Sick Children, Toronto, Ontario M5G 1X8, Canada.
Organizational Affiliation: