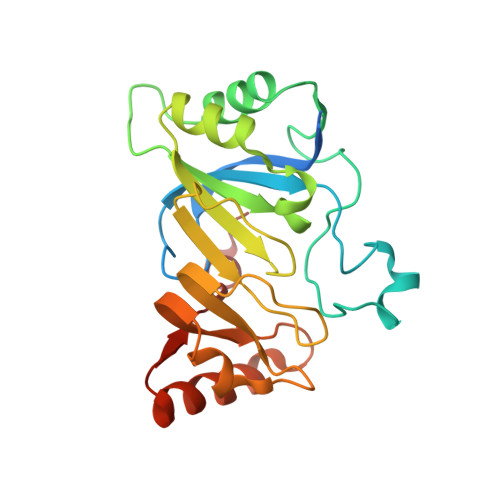
Radical SAM enzyme QueE defines a new minimal core fold and metal-dependent mechanism.
Dowling, D.P., Bruender, N.A., Young, A.P., McCarty, R.M., Bandarian, V., Drennan, C.L.(2014) Nat Chem Biol 10: 106-112
- PubMed: 24362703 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1426
- Primary Citation Related Structures:
4NJG, 4NJH, 4NJI, 4NJJ, 4NJK - PubMed Abstract:
7-carboxy-7-deazaguanine synthase (QueE) catalyzes a key S-adenosyl-L-methionine (AdoMet)- and Mg(2+)-dependent radical-mediated ring contraction step, which is common to the biosynthetic pathways of all deazapurine-containing compounds. QueE is a member of the AdoMet radical superfamily, which employs the 5'-deoxyadenosyl radical from reductive cleavage of AdoMet to initiate chemistry. To provide a mechanistic rationale for this elaborate transformation, we present the crystal structure of a QueE along with structures of pre- and post-turnover states. We find that substrate binds perpendicular to the [4Fe-4S]-bound AdoMet, exposing its C6 hydrogen atom for abstraction and generating the binding site for Mg(2+), which coordinates directly to the substrate. The Burkholderia multivorans structure reported here varies from all other previously characterized members of the AdoMet radical superfamily in that it contains a hypermodified (β6/α3) protein core and an expanded cluster-binding motif, CX14CX2C.
- 1] Howard Hughes Medical Institute, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA. [2] Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
Organizational Affiliation: