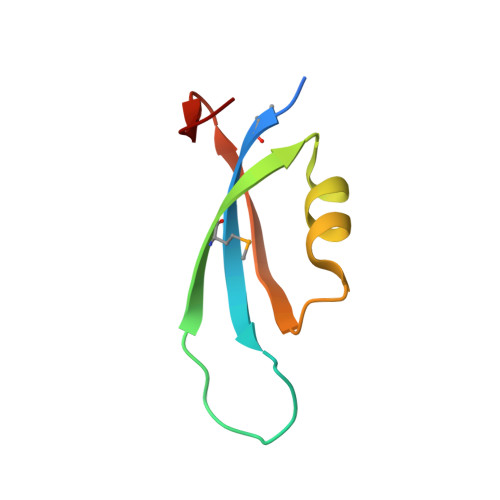
epsilon, a New Subunit of RNA Polymerase Found in Gram-Positive Bacteria
Keller, A.N., Yang, X., Wiedermannova, J., Delumeau, O., Krasny, L., Lewis, P.J.(2014) J Bacteriol 196: 3622-3632
- PubMed: 25092033
- DOI: https://doi.org/10.1128/JB.02020-14
- Primary Citation Related Structures:
4NJC - PubMed Abstract:
RNA polymerase in bacteria is a multisubunit protein complex that is essential for gene expression. We have identified a new subunit of RNA polymerase present in the high-A+T Firmicutes phylum of Gram-positive bacteria and have named it ε. Previously ε had been identified as a small protein (ω1) that copurified with RNA polymerase. We have solved the structure of ε by X-ray crystallography and show that it is not an ω subunit. Rather, ε bears remarkable similarity to the Gp2 family of phage proteins involved in the inhibition of host cell transcription following infection. Deletion of ε shows no phenotype and has no effect on the transcriptional profile of the cell. Determination of the location of ε within the assembly of RNA polymerase core by single-particle analysis suggests that it binds toward the downstream side of the DNA binding cleft. Due to the structural similarity of ε with Gp2 and the fact they bind similar regions of RNA polymerase, we hypothesize that ε may serve a role in protection from phage infection.
- School of Environmental and Life Sciences, University of Newcastle, Callghan, New South Wales, Australia.
Organizational Affiliation: