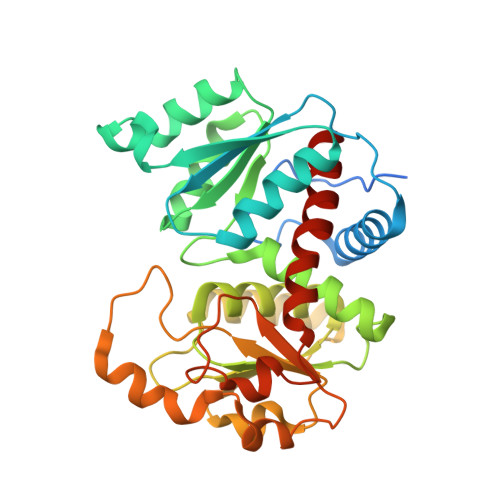
Crystal structures and kinetic properties of anabolic ornithine carbamoyltransferase from human pathogens Vibrio vulnificus and Bacillus anthracis
Shabalin, I.G., Handing, K., Cymborowski, M.T., Stam, J., Winsor, J., Shuvalova, L., Anderson, W.F., Minor, W.To be published.