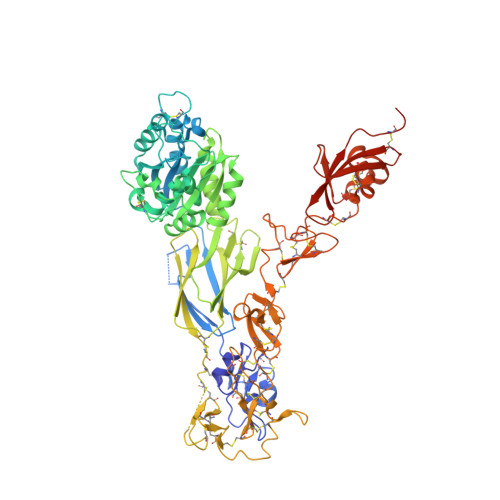
An internal ligand-bound, metastable state of a leukocyte integrin, alpha X beta 2.
Sen, M., Yuki, K., Springer, T.A.(2013) J Cell Biol 203: 629-642
- PubMed: 24385486 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1083/jcb.201308083
- Primary Citation Related Structures:
4NEH, 4NEN - PubMed Abstract:
How is massive conformational change in integrins achieved on a rapid timescale? We report crystal structures of a metastable, putative transition state of integrin αXβ2. The αXβ2 ectodomain is bent; however, a lattice contact stabilizes its ligand-binding αI domain in a high affinity, open conformation. Much of the αI α7 helix unwinds, loses contact with the αI domain, and reshapes to form an internal ligand that binds to the interface between the β propeller and βI domains. Lift-off of the αI domain above this platform enables a range of extensional and rotational motions without precedent in allosteric machines. Movements of secondary structure elements in the β2 βI domain occur in an order different than in β3 integrins, showing that integrin β subunits can be specialized to assume different intermediate states between closed and open. Mutations demonstrate that the structure trapped here is metastable and can enable rapid equilibration between bent and extended-open integrin conformations and up-regulation of leukocyte adhesiveness.
- Program in Cellular and Molecular Medicine, 2 Department of Medicine, 3 Department of Anethesiology, 4 Children's Hospital Boston, and 5 Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115.
Organizational Affiliation: