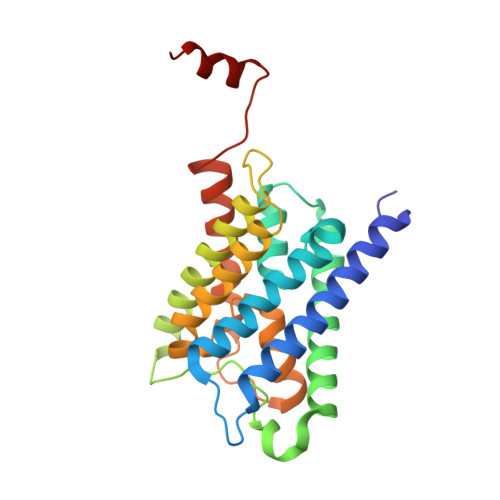
X-ray structure of human aquaporin 2 and its implications for nephrogenic diabetes insipidus and trafficking
Frick, A., Eriksson, U.K., de Mattia, F., Oberg, F., Hedfalk, K., Neutze, R., de Grip, W.J., Deen, P.M., Tornroth-Horsefield, S.(2014) Proc Natl Acad Sci U S A 111: 6305-6310
- PubMed: 24733887 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1321406111
- Primary Citation Related Structures:
4NEF - PubMed Abstract:
Human aquaporin 2 (AQP2) is a water channel found in the kidney collecting duct, where it plays a key role in concentrating urine. Water reabsorption is regulated by AQP2 trafficking between intracellular storage vesicles and the apical membrane. This process is tightly controlled by the pituitary hormone arginine vasopressin and defective trafficking results in nephrogenic diabetes insipidus (NDI). Here we present the X-ray structure of human AQP2 at 2.75 Å resolution. The C terminus of AQP2 displays multiple conformations with the C-terminal α-helix of one protomer interacting with the cytoplasmic surface of a symmetry-related AQP2 molecule, suggesting potential protein-protein interactions involved in cellular sorting of AQP2. Two Cd(2+)-ion binding sites are observed within the AQP2 tetramer, inducing a rearrangement of loop D, which facilitates this interaction. The locations of several NDI-causing mutations can be observed in the AQP2 structure, primarily situated within transmembrane domains and the majority of which cause misfolding and ER retention. These observations provide a framework for understanding why mutations in AQP2 cause NDI as well as structural insights into AQP2 interactions that may govern its trafficking.
- Department of Chemistry and Molecular Biology, University of Gothenburg, 405 30 Gothenburg, Sweden.
Organizational Affiliation: