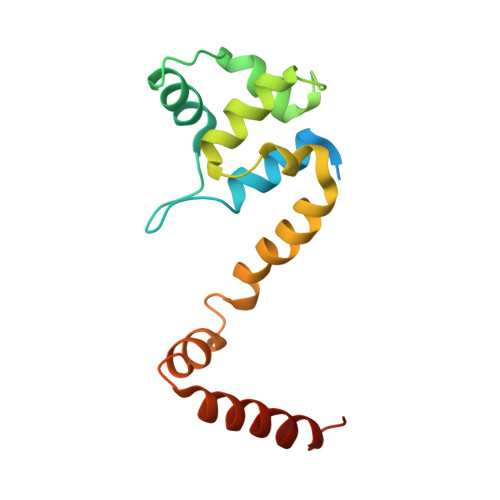
Comparison of four different crystal forms of the Mycobacterium tuberculosis ESX-1 secreted protein regulator EspR
Gangwar, S.P., Meena, S.R., Saxena, A.K.(2014) Acta Crystallogr Sect F Struct Biol Cryst Commun 70: 433-437
- PubMed: 24699733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14004166
- Primary Citation Related Structures:
4NDW - PubMed Abstract:
The Mycobacterium tuberculosis ESX-1 secreted protein regulator (EspR, Rv3849) is the key protein that delivers bacterial proteins into the host cell during mycobacterial infection. EspR binds directly to the espACD operon and is involved in transcriptional activation. In the current study, M. tuberculosis EspR has been crystallized and its X-ray structure has been determined at 3.3 Å resolution in a P3221 crystal form. EspR forms a physiological dimer in the crystal. Each EspR monomer contains an N-terminal helix-turn-helix DNA-binding domain and a C-terminal dimerization domain. The EspR structure in the P3221 crystal form was compared with previously determined EspR structures in P32, P21 and P212121 crystal forms. Structural comparison analysis indicated that the N-terminal helix-turn-helix domain of EspR acquires a rigid structure in the four crystal forms. However, significant structural differences were observed in the C-terminal domain of EspR in the P21 crystal form when compared with the P3221 and P32 crystal forms. The interaction, stabilization energy and buried surface area analysis of EspR in the four different crystal forms have provided information about the physiological dimer interface of EspR.
- School of Life Sciences, Jawaharlal Nehru University, New Delhi 110 067, India.
Organizational Affiliation: