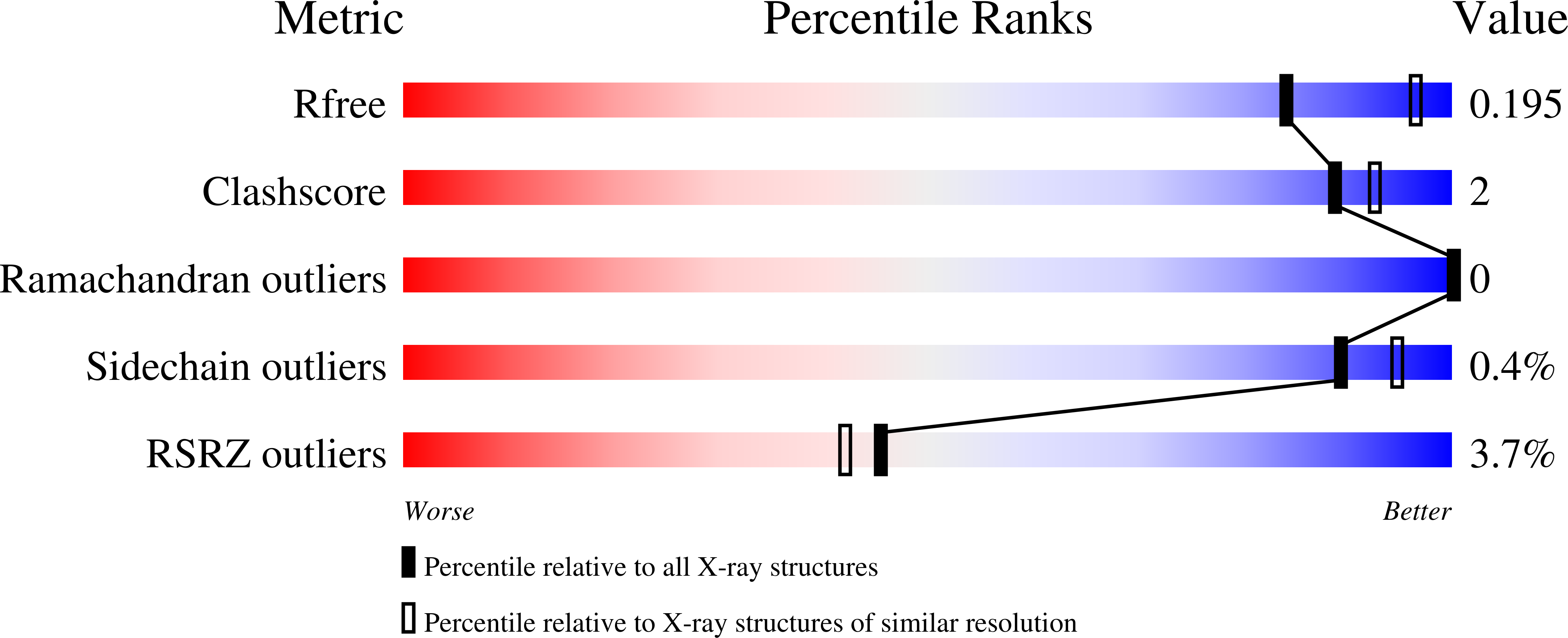
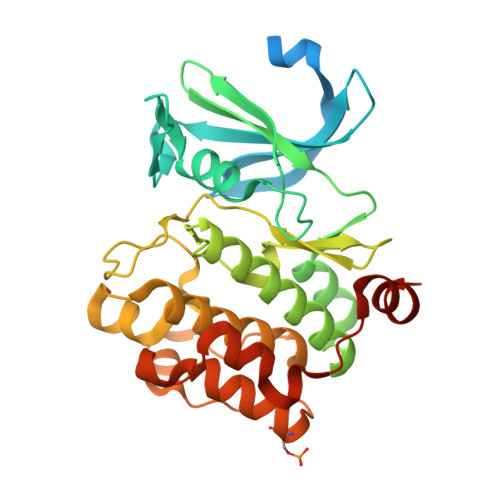
Structure Guided Optimization, in Vitro Activity, and in Vivo Activity of Pan-PIM Kinase Inhibitors.
Burger, M.T., Han, W., Lan, J., Nishiguchi, G., Bellamacina, C., Lindval, M., Atallah, G., Ding, Y., Mathur, M., McBride, C., Beans, E.L., Muller, K., Tamez, V., Zhang, Y., Huh, K., Feucht, P., Zavorotinskaya, T., Dai, Y., Holash, J., Castillo, J., Langowski, J., Wang, Y., Chen, M.Y., Garcia, P.D.(2013) ACS Med Chem Lett 4: 1193-1197
- PubMed: 24900629 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml400307j
- Primary Citation Related Structures:
4N6Y, 4N6Z, 4N70 - PubMed Abstract:
Proviral insertion of Moloney virus (PIM) 1, 2, and 3 kinases are serine/threonine kinases that normally function in survival and proliferation of hematopoietic cells. As high expression of PIM1, 2, and 3 is frequently observed in many human malignancies, including multiple myeloma, non-Hodgkins lymphoma, and myeloid leukemias, there is interest in determining whether selective PIM inhibition can improve outcomes of these human cancers. Herein, we describe our efforts toward this goal. The structure guided optimization of a singleton high throughput screening hit in which the potency against all three PIM isoforms was increased >10,000-fold to yield compounds with pan PIM K is < 10 pM, nanomolar cellular potency, and in vivo activity in an acute myeloid leukemia Pim-dependent tumor model is described.
- Global Discovery Chemistry/Oncology & Exploratory Chemistry, Novartis Institutes for Biomedical Research , 4560 Horton Street, Emeryville, California 94608, United States.
Organizational Affiliation: