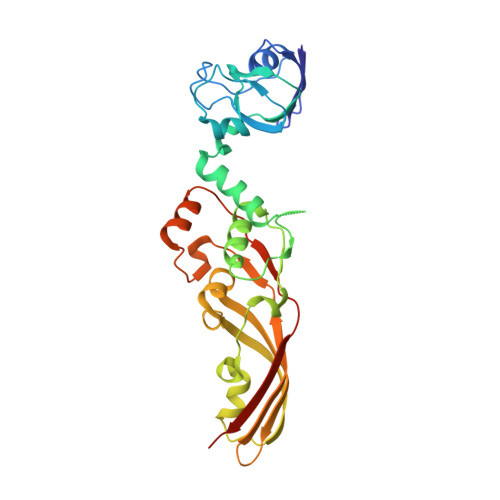
Structure of the atypical bacteriocin pectocin M2 implies a novel mechanism of protein uptake.
Grinter, R., Josts, I., Zeth, K., Roszak, A.W., McCaughey, L.C., Cogdell, R.J., Milner, J.J., Kelly, S.M., Byron, O., Walker, D.(2014) Mol Microbiol 93: 234-246
- PubMed: 24865810 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/mmi.12655
- Primary Citation Related Structures:
4N58, 4N59 - PubMed Abstract:
The colicin-like bacteriocins are potent protein antibiotics that have evolved to efficiently cross the outer membrane of Gram-negative bacteria by parasitizing nutrient uptake systems. We have structurally characterized the colicin M-like bacteriocin, pectocin M2, which is active against strains of Pectobacterium spp. This unusual bacteriocin lacks the intrinsically unstructured translocation domain that usually mediates translocation of these bacteriocins across the outer membrane, containing only a single globular ferredoxin domain connected to its cytotoxic domain by a flexible α-helix, which allows it to adopt two distinct conformations in solution. The ferredoxin domain of pectocin M2 is homologous to plant ferredoxins and allows pectocin M2 to parasitize a system utilized by Pectobacterium to obtain iron during infection of plants. Furthermore, we identify a novel ferredoxin-containing bacteriocin pectocin P, which possesses a cytotoxic domain homologous to lysozyme, illustrating that the ferredoxin domain acts as a generic delivery module for cytotoxic domains in Pectobacterium.
- Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow, G12 8QQ, UK.
Organizational Affiliation: