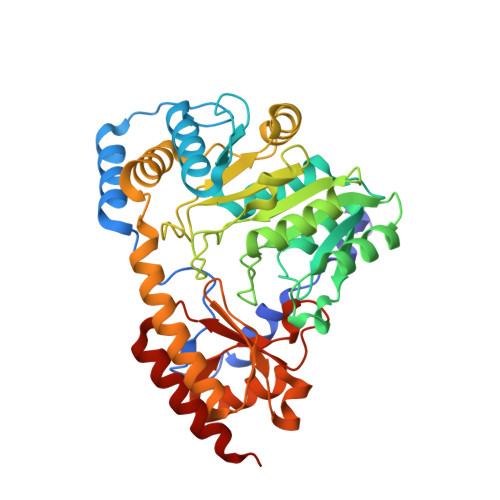
Crystal structure of the aromatic-amino-acid aminotransferase from Streptococcus mutans.
Cong, X., Li, X., Li, S.(2019) Acta Crystallogr F Struct Biol Commun 75: 141-146
- PubMed: 30713166
- DOI: https://doi.org/10.1107/S2053230X18018472
- Primary Citation Related Structures:
4MY5 - PubMed Abstract:
Streptococcus mutans, a facultatively aerobic and Gram-positive bacterium, is the primary causative agent of dental caries and contributes to the multispecies biofilm known as dental plaque. In this study, the aromatic-amino-acid aminotransferase from Streptococcus mutans (SmAroAT) was recombinantly expressed in Escherichia coli. An effective purification protocol was established. The recombinant protein was crystallized using the hanging-drop vapor-diffusion method with PEG 3350 as the primary precipitant. The crystal structure of SmAroAT was solved at 2.2 Å resolution by the molecular-replacement method. Structural analysis indicated that the proteins of the aromatic-amino-acid aminotransferase family have conserved structural elements that might play a role in substrate binding. These results may help in obtaining a better understanding of the catabolism and biosynthesis of aromatic amino acids.
- Central Laboratory, Capital Medical University, Beijing 100069, People's Republic of China.
Organizational Affiliation: