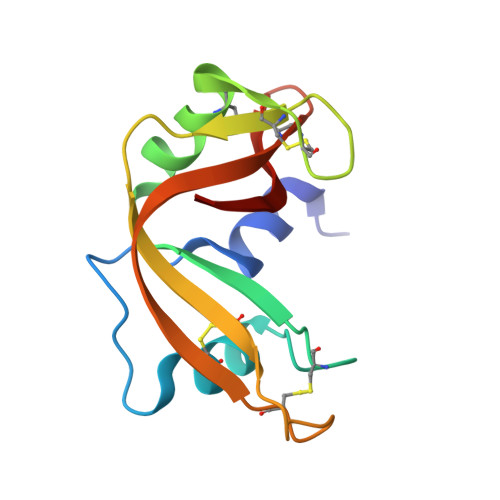
Interactions of gold-based drugs with proteins: crystal structure of the adduct formed between ribonuclease A and a cytotoxic gold(iii) compound.
Messori, L., Scaletti, F., Massai, L., Cinellu, M.A., Russo Krauss, I., di Martino, G., Vergara, A., Paduano, L., Merlino, A.(2014) Metallomics 6: 233-236
- PubMed: 24287583 Search on PubMed
- DOI: https://doi.org/10.1039/c3mt00265a
- Primary Citation Related Structures:
4MXF - PubMed Abstract:
The reaction of Auoxo6, a dinuclear gold(III) complex, with the model protein bovine pancreatic ribonuclease is explored here by X-ray diffraction and ESI mass spectrometry. Data provide clues on the processes of adduct formation and of enzyme inhibition and, inductively, on the likely mode of action of this metallodrug.
- Department of Chemistry "Ugo Schiff", Via della Lastruccia 3, 50019 Sesto Fiorentino (FI), Italy.
Organizational Affiliation: