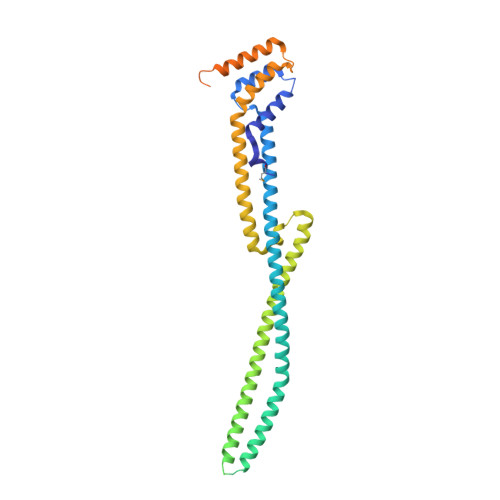
Structure of the N-terminal domain of the effector protein LegC3 from Legionella pneumophila.
Yao, D., Cherney, M., Cygler, M.(2014) Acta Crystallogr D Biol Crystallogr 70: 436-441
- PubMed: 24531477
- DOI: https://doi.org/10.1107/S139900471302991X
- Primary Citation Related Structures:
4MU6 - PubMed Abstract:
Legionella pneumophila secretes over 300 effectors during the invasion of human cells. The functions of only a small number of them have been identified. LegC3 is one of the identified effectors, which is believed to act by inhibiting vacuolar fusion. It contains two predicted transmembrane helices that divide the protein into a larger N-terminal domain and a smaller C-terminal domain. The function of LegC3 has been shown to be associated primarily with the N-terminal domain, which contains coiled-coil sequence motifs. The structure of the N-terminal domain has been determined and it is shown that it is highly α-helical and contains a helical bundle followed by a long antiparallel coiled-coil. No similar protein fold has been observed in the PDB. A long loop at the tip of the coiled-coil distal from the membrane is disordered and may be important for interaction with an as yet unidentified protein.
- Department of Biochemistry, University of Saskatchewan, 107 Wiggins Road, Saskatoon, SK S7N 5E5, Canada.
Organizational Affiliation: