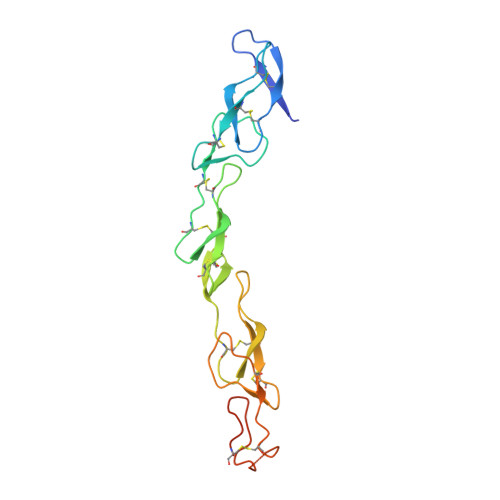
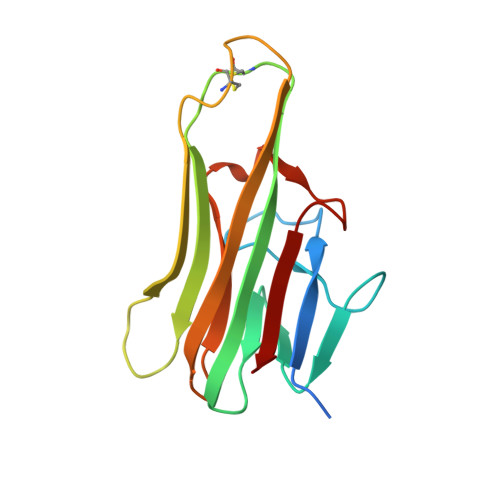
Crystal Structure of the Complex of Human FasL and Its Decoy Receptor DcR3.
Liu, W., Ramagopal, U., Cheng, H., Bonanno, J.B., Toro, R., Bhosle, R., Zhan, C., Almo, S.C.(2016) Structure 24: 2016-2023
- PubMed: 27806260 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2016.09.009
- Primary Citation Related Structures:
4MSV, 5L19, 5L36 - PubMed Abstract:
The apoptotic effect of FasL:Fas signaling is disrupted by DcR3, a unique secreted member of the tumor necrosis factor receptor superfamily, which also binds and neutralizes TL1A and LIGHT. DcR3 is highly elevated in patients with various tumors and contributes to mechanisms by which tumor cells to evade host immune surveillance. Here we report the crystal structure of FasL in complex with DcR3. Comparison of FasL:DcR3 structure with our earlier TL1A:DcR3 and LIGHT:DcR3 structures supports a paradigm involving the recognition of invariant main-chain and conserved side-chain functionalities, which is responsible for the recognition of multiple TNF ligands exhibited by DcR3. The FasL:DcR3 structure also provides insight into the FasL:Fas recognition surface. We demonstrate that the ability of recombinant FasL to induce Jurkat cell apoptosis is significantly enhanced by native glycosylation or by structure-inspired mutations, both of which result in reduced tendency to aggregate. All of these activities are efficiently inhibited by recombinant DcR3.
- Department of Biochemistry, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA; Department of Microbiology and Immunology, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA.
Organizational Affiliation: