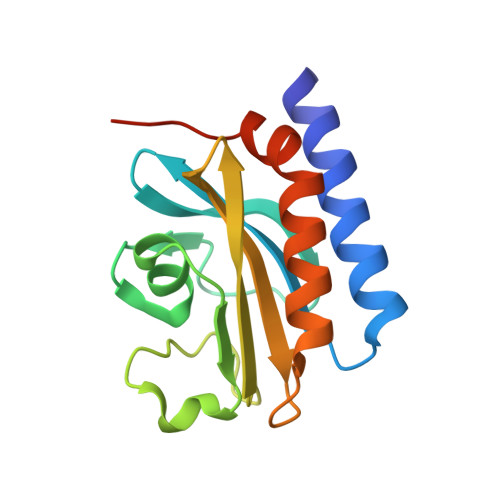
Structural and biochemical analysis of a type II free methionine-R-sulfoxide reductase from Thermoplasma acidophilum
Kim, H.S., Kwak, G.H., Lee, K., Jo, C.H., Hwang, K.Y., Kim, H.Y.(2014) Arch Biochem Biophys 560: 10-19
- PubMed: 25043974 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2014.07.009
- Primary Citation Related Structures:
4MMN, 4MN7 - PubMed Abstract:
Free methionine-R-sulfoxide reductase (fRMsr) enzymes only reduce the free form of methionine-R-sulfoxide and can be grouped into two types with respect to the number of conserved Cys residues in the active sites. In this work, the crystal structures of type II fRMsr from Thermoplasma acidophilum (TafRMsr), which contains two conserved Cys, have been determined in native form and in a complex with the substrate. The overall structure of TafRMsr consists of a central β-sheet encompassed by a two-α-helix bundle flanking on one side and one small α-helix on the other side. Based on biochemical and growth complementation assays, Cys(84) is demonstrated to be the catalytic Cys. The data also show that TafRMsr functions as an antioxidant protein. Structural analyses reveal insights into substrate recognition and orientation, conformational changes in the active site during substrate binding, and the role of active site residues in substrate binding. A model for the catalytic mechanism of type II TafRMsr is suggested, in which intramolecular disulfide bond formation is not involved. In addition, the biochemical, enzymatic, and structural properties of type II TafRMsr are compared with those of type I enzymes.
- Department of Biosystems and Biotechnology, College of Life Sciences and Biotechnology, Korea University, Seoul 136-701, Republic of Korea.
Organizational Affiliation: