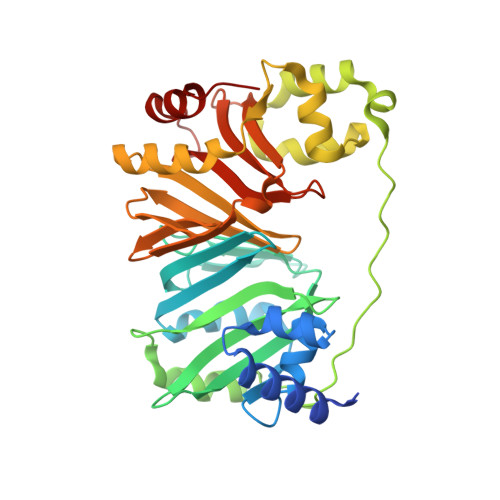
Structural analysis and identification of PhuS as a heme-degrading enzyme from Pseudomonas aeruginosa.
Lee, M.J., Schep, D., McLaughlin, B., Kaufmann, M., Jia, Z.(2014) J Mol Biology 426: 1936-1946
- PubMed: 24560694 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2014.02.013
- Primary Citation Related Structures:
4MF9, 4MGF - PubMed Abstract:
Bacterial pathogens require iron for proliferation and pathogenesis. Pseudomonas aeruginosa is a prevalent Gram-negative opportunistic human pathogen that takes advantage of immunocompromised hosts and encodes a number of proteins for uptake and utilization of iron. Here we report the crystal structures of PhuS, previously known as the cytoplasmic heme-trafficking protein from P. aeruginosa, in both the apo- and the holo-forms. In comparison to its homologue ChuS from Escherichia coli O157:H7, the heme orientation is rotated 180° across the α-γ axis, which may account for some of the unique functional properties of PhuS. In contrast to previous findings, heme binding does not result in an overall conformational change of PhuS. We employed spectroscopic analysis and CO measurement by gas chromatography to analyze heme degradation, demonstrating that PhuS is capable of degrading heme using ascorbic acid or cytochrome P450 reductase-NADPH as an electron donor and produces five times more CO than ChuS. Addition of catalase slows down but does not stop PhuS-catalyzed heme degradation. Through spectroscopic and mass spectrometry analysis, we identified the enzymatic product of heme degradation to be verdoheme. These data taken together suggest that PhuS is a potent heme-degrading enzyme, in addition to its proposed heme-trafficking function.
- Department of Biomedical and Molecular Sciences, Queen's University, Kingston, ON, Canada K7L 3N6.
Organizational Affiliation: