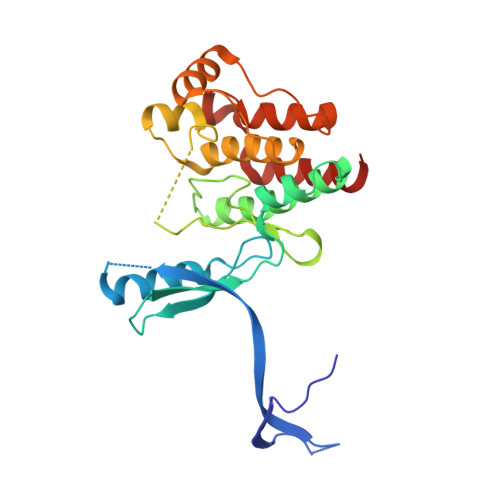
Structure-based design and synthesis of potent benzothiazole inhibitors of interleukin-2 inducible T cell kinase (ITK).
Mackinnon, C.H., Lau, K., Burch, J.D., Chen, Y., Dines, J., Ding, X., Eigenbrot, C., Heifetz, A., Jaochico, A., Johnson, A., Kraemer, J., Kruger, S., Krulle, T.M., Liimatta, M., Ly, J., Maghames, R., Montalbetti, C.A., Ortwine, D.F., Perez-Fuertes, Y., Shia, S., Stein, D.B., Trani, G., Vaidya, D.G., Wang, X., Bromidge, S.M., Wu, L.C., Pei, Z.(2013) Bioorg Med Chem Lett 23: 6331-6335
- PubMed: 24138940 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2013.09.069
- Primary Citation Related Structures:
4MF0, 4MF1 - PubMed Abstract:
Inhibition of the non-receptor tyrosine kinase ITK, a component of the T-cell receptor signalling cascade, may represent a novel treatment for allergic asthma. Here we report the structure-based optimization of a series of benzothiazole amides that demonstrate sub-nanomolar inhibitory potency against ITK with good cellular activity and kinase selectivity. We also elucidate the binding mode of these inhibitors by solving the X-ray crystal structures of several inhibitor-ITK complexes.
- Evotec (UK) Ltd, 114 Milton Park, Abingdon, Oxfordshire OX14 4SA, United Kingdom.
Organizational Affiliation: