Discovery of N-(benzo[1,2,3]triazol-1-yl)-N-(benzyl)acetamido)phenyl) carboxamides as severe acute respiratory syndrome coronavirus (SARS-CoV) 3CLpro inhibitors: Identification of ML300 and noncovalent nanomolar inhibitors with an induced-fit binding.
Turlington, M., Chun, A., Tomar, S., Eggler, A., Grum-Tokars, V., Jacobs, J., Daniels, J.S., Dawson, E., Saldanha, A., Chase, P., Baez-Santos, Y.M., Lindsley, C.W., Hodder, P., Mesecar, A.D., Stauffer, S.R.(2013) Bioorg Med Chem Lett 23: 6172-6177
- PubMed: 24080461 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2013.08.112
- Primary Citation Related Structures:
4MDS - PubMed Abstract:
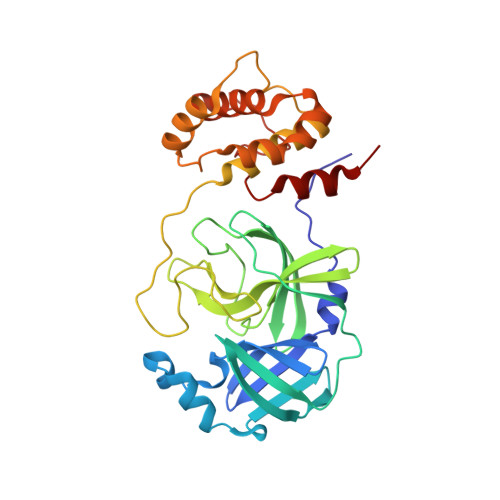
Herein we report the discovery and SAR of a novel series of SARS-CoV 3CLpro inhibitors identified through the NIH Molecular Libraries Probe Production Centers Network (MLPCN). In addition to ML188, ML300 represents the second probe declared for 3CLpro from this collaborative effort. The X-ray structure of SARS-CoV 3CLpro bound with a ML300 analog highlights a unique induced-fit reorganization of the S2-S4 binding pockets leading to the first sub-micromolar noncovalent 3CLpro inhibitors retaining a single amide bond.
- Department of Pharmacology, Vanderbilt University Medical Center, Nashville, TN 37232, USA; Vanderbilt Center for Neuroscience Drug Discovery, Vanderbilt University Medical Center, Nashville, TN 37232, USA; Vanderbilt Specialized Chemistry Center for Probe Development (MLPCN), Nashville, TN 37232, USA.
Organizational Affiliation: