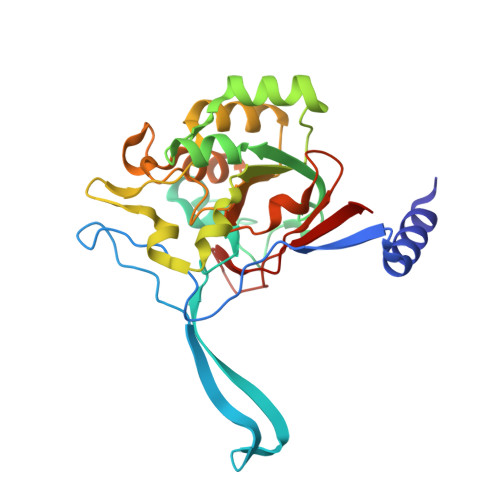
A proton wire and water channel revealed in the crystal structure of isatin hydrolase.
Bjerregaard-Andersen, K., Sommer, T., Jensen, J.K., Jochimsen, B., Etzerodt, M., Morth, J.P.(2014) J Biological Chem 289: 21351-21359
- PubMed: 24917679 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.568824
- Primary Citation Related Structures:
4J0N, 4M8D - PubMed Abstract:
The high resolution crystal structures of isatin hydrolase from Labrenzia aggregata in the apo and the product state are described. These are the first structures of a functionally characterized metal-dependent hydrolase of this fold. Isatin hydrolase converts isatin to isatinate and belongs to a novel family of metalloenzymes that include the bacterial kynurenine formamidase. The product state, mimicked by bound thioisatinate, reveals a water molecule that bridges the thioisatinate to a proton wire in an adjacent water channel and thus allows the proton released by the reaction to escape only when the product is formed. The functional proton wire present in isatin hydrolase isoform b represents a unique catalytic feature common to all hydrolases is here trapped and visualized for the first time. The local molecular environment required to coordinate thioisatinate allows stronger and more confident identification of orthologous genes encoding isatin hydrolases within the prokaryotic kingdom. The isatin hydrolase orthologues found in human gut bacteria raise the question as to whether the indole-3-acetic acid degradation pathway is present in human gut flora.
- From the Norwegian Center of Molecular Medicine, Nordic EMBL Partnership University of Oslo, Gaustadalléen 21, 0349 Oslo, Norway, the Department for Molecular Biology and Genetics, Aarhus University, Gustav Wieds vej 10C, DK-8000 Aarhus, Denmark.
Organizational Affiliation: