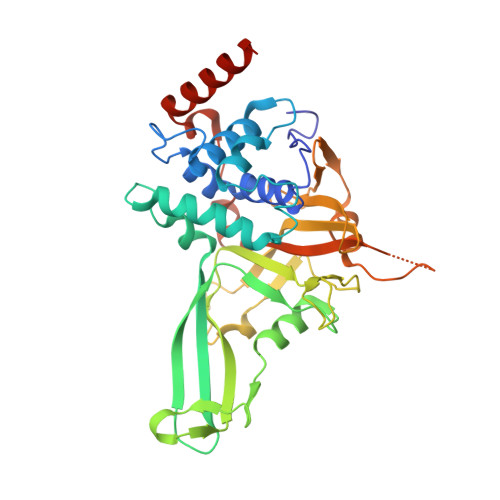
A 2.2 angstrom resolution structure of the USP7 catalytic domain in a new space group elaborates upon structural rearrangements resulting from ubiquitin binding.
Molland, K., Zhou, Q., Mesecar, A.D.(2014) Acta Crystallogr Sect F Struct Biol Cryst Commun 70: 283-287
- PubMed: 24598911 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14002519
- Primary Citation Related Structures:
4M5W, 4M5X - PubMed Abstract:
A sparse-matrix screen for new crystallization conditions for the USP7 catalytic domain (USP7CD) led to the identification of a condition in which crystals grow reproducibly in 24-48 h. Variation of the halide metal, growth temperature and seed-stock concentration resulted in a shift in space group from P21 with two molecules in the asymmetric unit to C2 with one molecule in the asymmetric unit. Representative structures from each space group were determined to 2.2 Å resolution and these structures support previous findings that the catalytic triad and switching loop are likely to be in unproductive conformations in the absence of ubiquitin (Ub). Importantly, the new structures reveal previously unobserved electron density for blocking loop 1 (BL1) residues 410-419. The new structures indicate a distinct rearrangement of the USP7 BL1 compared with its position in the presence of bound Ub.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.
Organizational Affiliation: