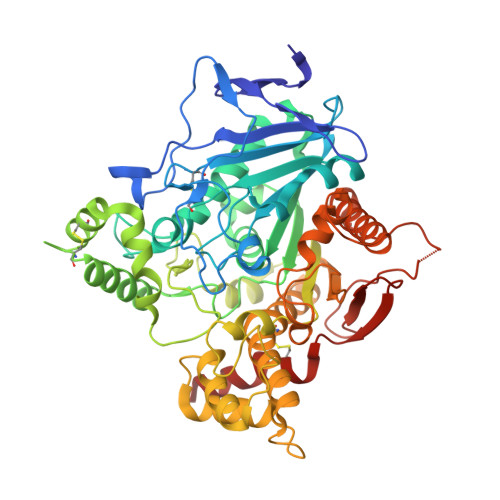
Structures of human acetylcholinesterase bound to dihydrotanshinone I and territrem B show peripheral site flexibility.
Cheung, J., Gary, E.N., Shiomi, K., Rosenberry, T.L.(2013) ACS Med Chem Lett 4: 1091-1096
- PubMed: 24900610 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml400304w
- Primary Citation Related Structures:
4M0E, 4M0F - PubMed Abstract:
Acetylcholinesterase is a critical enzyme that regulates neurotransmission by degrading the neurotransmitter acetylcholine in synapses of the nervous system. It is an important target for both therapeutic drugs that treat Alzheimer's disease and chemical warfare agents that cripple the nervous system and cause death through paralysis. The enzyme has both catalytic and peripheral sites to which inhibitors may bind. Structures of recombinant human acetylcholinesterase in complex with the natural product inhibitors dihydrotanshinone I and territrem B reveal dihydrotanshinone I binding that is specific to only the peripheral site and territrem B binding that spans both sites and distorts the protein backbone in the peripheral site. These inhibitors may function as important molecular templates for therapeutics used for treatment of disease and protection against nerve agents.
- New York Structural Biology Center, New York, New York 10027, United States.
Organizational Affiliation: