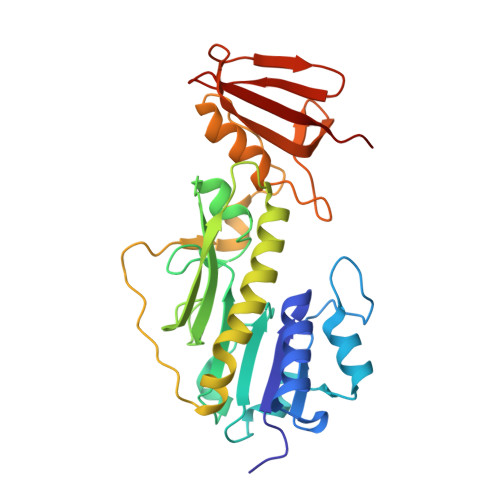
Structure of a sugar N-formyltransferase from Campylobacter jejuni.
Thoden, J.B., Goneau, M.F., Gilbert, M., Holden, H.M.(2013) Biochemistry 52: 6114-6126
- PubMed: 23898784
- DOI: https://doi.org/10.1021/bi4009006
- Primary Citation of Related Structures:
4LXQ, 4LXT, 4LXU, 4LXX, 4LXY, 4LY0, 4LY3 - PubMed Abstract:
The O-antigens, which are components of the outer membranes of Gram-negative bacteria, are responsible for the wide species variations seen in nature and are thought to play a role in bacterial virulence. They often contain unusual dideoxysugars such as 3,6-dideoxy-3-formamido-d-glucose (Qui3NFo). Here, we describe a structural and functional investigation of the protein C8J_1081 from Campylobacter jejuni 81116, which is involved in the biosynthesis of Qui3NFo. Specifically, the enzyme, hereafter referred to as WlaRD, catalyzes the N-formylation of dTDP-3,6-dideoxy-3-amino-d-glucose (dTDP-Qui3N) using N(10)-formyltetrahydrofolate as the carbon source. For this investigation, seven X-ray structures of WlaRD, in complexes with various dTDP-linked sugars and cofactors, were determined to resolutions of 1.9 Å or better. One of the models, with bound N(10)-formyltetrahydrofolate and dTDP, represents the first glimpse of an N-formyltransferase with its natural cofactor. Another model contains the reaction products, tetrahydrofolate and dTDP-Qui3NFo. In combination, the structures provide snapshots of the WlaRD active site before and after catalysis. On the basis of these structures, three amino acid residues were targeted for study: Asn 94, His 96, and Asp 132. Mutations of any of these residues resulted in a complete loss of enzymatic activity. Given the position of His 96 in the active site, it can be postulated that it functions as the active site base to remove a proton from the sugar amino group as it attacks the carbonyl carbon of the N-10 formyl group of the cofactor. Enzyme assays demonstrate that WlaRD is also capable of utilizing dTDP-3,6-dideoxy-3-amino-d-galactose (dTDP-Fuc3N) as a substrate, albeit at a much reduced catalytic efficiency.
- Department of Biochemistry, University of Wisconsin , Madison, Wisconsin 53706, United States.
Organizational Affiliation: