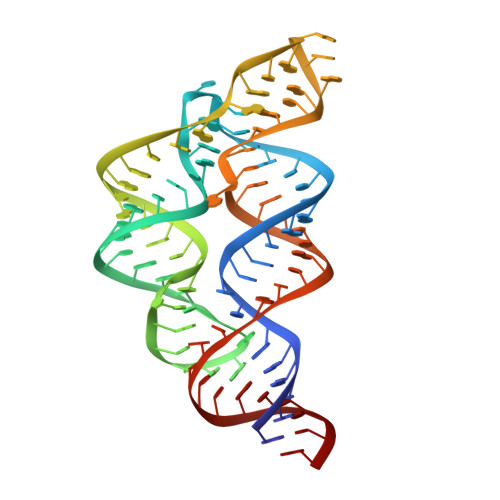
A Disconnect between High-Affinity Binding and Efficient Regulation by Antifolates and Purines in the Tetrahydrofolate Riboswitch.
Trausch, J.J., Batey, R.T.(2014) Chem Biol 21: 205-216
- PubMed: 24388757 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2013.11.012
- Primary Citation Related Structures:
4LVV, 4LVW, 4LVX, 4LVY, 4LVZ, 4LW0 - PubMed Abstract:
The tetrahydrofolate (THF) riboswitch regulates folate transport and metabolism in a number of Firmicutes by cooperatively binding two molecules of THF. To further understand this riboswitch's specificity for THF, binding and regulatory activity of a series of THF analogs and antifolates were examined. Our data reveal that although binding is dominated by the RNA's interactions with the pterin moiety, the para-aminobenzoic acid (pABA) moiety plays a significant role in transcriptional regulation. Further, we find that adenine and several other analogs bind with high affinity by an alternative binding mechanism. Despite a similar affinity to THF, adenine is a poor regulator of transcriptional attenuation. These results demonstrate that binding alone does not determine a compound's effectiveness in regulating the activity of the riboswitch-a complication in current efforts to develop antimicrobials that target these RNAs.
- Department of Chemistry and Biochemistry, University of Colorado at Boulder, Campus Box 596, Boulder, CO 80309-0596, USA.
Organizational Affiliation: