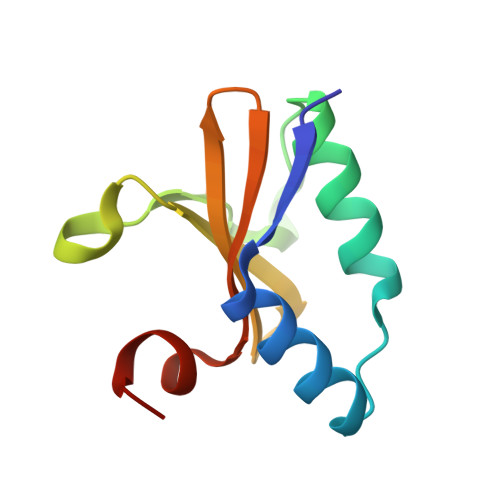
Crystal structure of apo and copper bound HP0894 toxin from Helicobacter pylori 26695 and insight into mRNase activity
Pathak, C., Im, H., Yang, Y.J., Yoon, H.J., Kim, H.M., Kwon, A.R., Lee, B.J.(2013) Biochim Biophys Acta 1834: 2579-2590
- PubMed: 24060809 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2013.09.006
- Primary Citation Related Structures:
4LS4, 4LSY, 4LTT - PubMed Abstract:
The toxin-antitoxin (TA) systems widely spread among bacteria and archaea are important for antibiotic resistance and microorganism virulence. The bacterial kingdom uses TA systems to adjust the global level of gene expression and translation through RNA degradation. In Helicobacter pylori, only two TA systems are known thus far. Our previous studies showed that HP0894-HP0895 acts as a TA system and that HP0894 exhibits intrinsic RNase activity. However, the precise molecular basis for interaction with substrate or antitoxin and the mechanism of mRNA cleavage remain unclear. Therefore, in an attempt to shed some light on the mechanism behind the TA system of HP0894-HP0895, here we present the crystal structures of apo- and metal-bound H. pylori 0894 at 1.28Å and 1.89Å, respectively. Through the combined approach of structural analysis and structural homology search, the amino acids involved in mRNase active site were monitored and the reorientations of different residues were discussed in detail. In the mRNase active site of HP0894 toxin, His84 acts as a catalytic residue and reorients itself to exhibit this type of activity, acting as a general acid in an acid-base catalysis reaction, while His47 and His60 stabilize the transition state. Lys52, Glu58, Asp64 and Arg80 have phosphate binding and specific sequence recognition. Glu58 also acts as a general base, and substrate reorientation is caused by Phe88. Based on experimental findings, a model for antitoxin binding could be suggested.
- Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Seoul 151-742, Republic of Korea.
Organizational Affiliation: