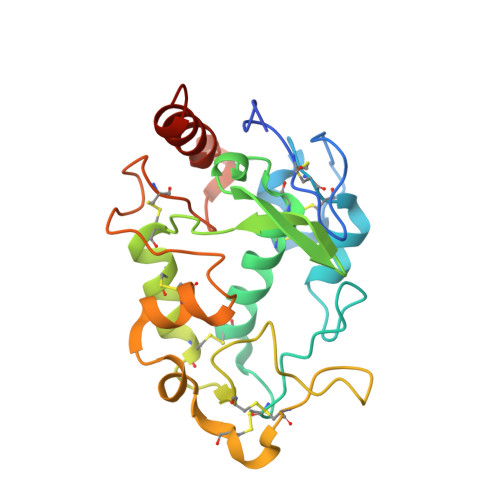
Structural basis for molecular recognition of folic acid by folate receptors.
Chen, C., Ke, J., Zhou, X.E., Yi, W., Brunzelle, J.S., Li, J., Yong, E.L., Xu, H.E., Melcher, K.(2013) Nature 500: 486-489
- PubMed: 23851396
- DOI: https://doi.org/10.1038/nature12327
- Primary Citation Related Structures:
4LRH - PubMed Abstract:
Folate receptors (FRα, FRβ and FRγ) are cysteine-rich cell-surface glycoproteins that bind folate with high affinity to mediate cellular uptake of folate. Although expressed at very low levels in most tissues, folate receptors, especially FRα, are expressed at high levels in numerous cancers to meet the folate demand of rapidly dividing cells under low folate conditions. The folate dependency of many tumours has been therapeutically and diagnostically exploited by administration of anti-FRα antibodies, high-affinity antifolates, folate-based imaging agents and folate-conjugated drugs and toxins. To understand how folate binds its receptors, we determined the crystal structure of human FRα in complex with folic acid at 2.8 Å resolution. FRα has a globular structure stabilized by eight disulphide bonds and contains a deep open folate-binding pocket comprised of residues that are conserved in all receptor subtypes. The folate pteroate moiety is buried inside the receptor, whereas its glutamate moiety is solvent-exposed and sticks out of the pocket entrance, allowing it to be conjugated to drugs without adversely affecting FRα binding. The extensive interactions between the receptor and ligand readily explain the high folate-binding affinity of folate receptors and provide a template for designing more specific drugs targeting the folate receptor system.
- Program for Structural Biology and Drug Discovery, Van Andel Research Institute, 333 Bostwick Avenue North East, Grand Rapids, Michigan 49503, USA.
Organizational Affiliation: