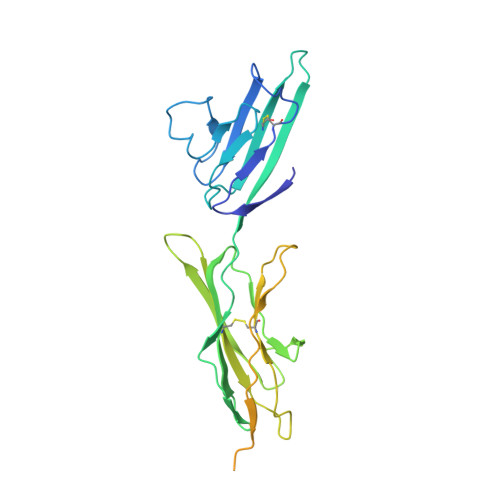
Structural insights into the oligomerization mode of the human receptor for advanced glycation end-products.
Yatime, L., Andersen, G.R.(2013) FEBS J 280: 6556-6568
- PubMed: 24119142 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12556
- Primary Citation Related Structures:
4LP4, 4LP5 - PubMed Abstract:
The receptor for advanced glycation end-products (RAGE) is a pattern recognition receptor sensing endogenous stress signals associated with the development of various diseases, including diabetes, vascular complications, Alzheimer's disease and cancer. RAGE ligands include advanced glycation end-products, S100 proteins, high mobility group box 1 protein and amyloid β-peptides/fibrils. Their signalling through RAGE induces a sustained inflammation that accentuates tissue damage, thereby participating in disease progression. Receptor oligomerization appears to be a crucial parameter for the formation of active signalling complexes, although the precise mode of oligomerization remains unclear in the context of these various ligands. In the present study, we report the first crystal structure of the VC1C2 fragment of the RAGE ectodomain. This structure provides the first description of the C2 domain in the context of the entire ectodomain and supports the observation of its conformational freedom relative to the rigid VC1 domain tandem. In addition, we have obtained a new crystal structure of the RAGE VC1 fragment. The packing in both crystal structures reveals an association of the RAGE molecules through contacts between two V domains and the physiological relevance of this homodimerization mode is discussed. Based on homology with single-pass transmembrane receptors, we also suggest RAGE dimerization through a conserved GxxxG motif within its transmembrane domain. A multimodal homodimerization strategy of RAGE is proposed to form the structural basis for ligand-specific complex formation and signalling functions, as well as for RAGE-mediated cell adhesion. hRAGE_VC1C2 and hRAGE_VC1C2 bind by x-ray crystallography (View interaction) hRAGE_VC1 and hRAGE_VC1 bind by x-ray crystallography (View interaction).
- Department of Molecular Biology and Genetics, Aarhus University, Denmark.
Organizational Affiliation: