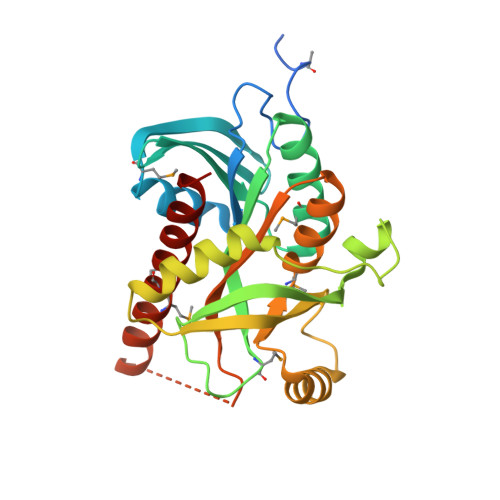
Crystal structure of uridine phosphorylase from Vibrio fischeri ES114, NYSGRC Target 29520.
Malashkevich, V.N., Bonanno, J.B., Bhosle, R., Toro, R., Hillerich, B., Gizzi, A., Garforth, S., Kar, A., Chan, M.K., Lafluer, J., Patel, H., Matikainen, B., Chamala, S., Lim, S., Celikgil, A., Villegas, G., Evans, B., Love, J., Fiser, A., Khafizov, K., Seidel, R., Almo, S.C.To be published.