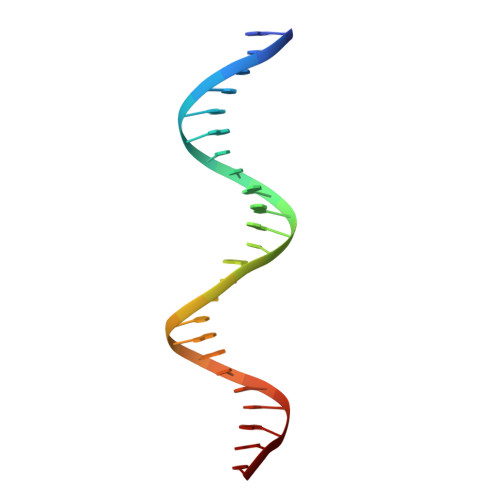
Structural mechanism of transcription regulation of the Staphylococcus aureus multidrug efflux operon mepRA by the MarR family repressor MepR.
Birukou, I., Seo, S.M., Schindler, B.D., Kaatz, G.W., Brennan, R.G.(2014) Nucleic Acids Res 42: 2774-2788
- PubMed: 24293644 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkt1215
- Primary Citation Related Structures:
4LLL, 4LLN - PubMed Abstract:
The multidrug efflux pump MepA is a major contributor to multidrug resistance in Staphylococcus aureus. MepR, a member of the multiple antibiotic resistance regulator (MarR) family, represses mepA and its own gene. Here, we report the structure of a MepR-mepR operator complex. Structural comparison of DNA-bound MepR with 'induced' apoMepR reveals the large conformational changes needed to allow the DNA-binding winged helix-turn-helix motifs to interact with the consecutive major and minor grooves of the GTTAG signature sequence. Intriguingly, MepR makes no hydrogen bonds to major groove nucleobases. Rather, recognition-helix residues Thr60, Gly61, Pro62 and Thr63 make sequence-specifying van der Waals contacts with the TTAG bases. Removing these contacts dramatically affects MepR-DNA binding activity. The wings insert into the flanking minor grooves, whereby residue Arg87, buttressed by Asp85, interacts with the O2 of T4 and O4' ribosyl oxygens of A23 and T4. Mutating Asp85 and Arg87, both conserved throughout the MarR family, markedly affects MepR repressor activity. The His14':Arg59 and Arg10':His35:Phe108 interaction networks stabilize the DNA-binding conformation of MepR thereby contributing significantly to its high affinity binding. A structure-guided model of the MepR-mepA operator complex suggests that MepR dimers do not interact directly and cooperative binding is likely achieved by DNA-mediated allosteric effects.
- Department of Biochemistry, Duke University School of Medicine, 307 Research Drive, Durham, NC 27710, USA, The John D. Dingell Department of Veterans Affairs Medical Center, B4333 JD Dingel VA Medical Center, 4646 John R, Detroit, MI 48201, USA and Department of Internal Medicine, Division of Infectious Diseases, Wayne State University School of Medicine, 5 Hudson, Harper University Hospital, 3990 John R, Detroit, MI 48201, USA.
Organizational Affiliation: