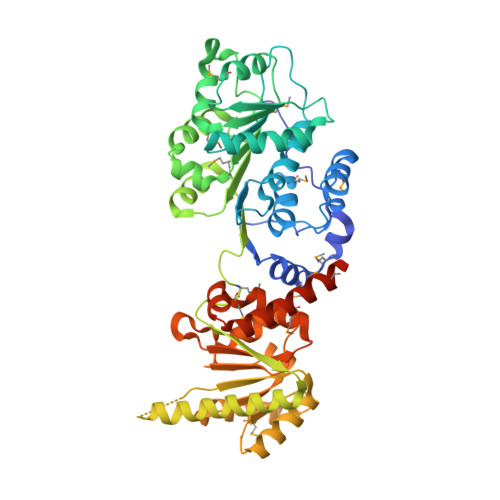
Crystal structure of Prp5p reveals interdomain interactions that impact spliceosome assembly.
Zhang, Z.M., Yang, F., Zhang, J., Tang, Q., Li, J., Gu, J., Zhou, J., Xu, Y.Z.(2013) Cell Rep 5: 1269-1278
- PubMed: 24290758
- DOI: https://doi.org/10.1016/j.celrep.2013.10.047
- Primary Citation Related Structures:
4LJY, 4LK2 - PubMed Abstract:
The DEAD-box adenosine triphosphatase (ATPase) Prp5p facilitates U2 small nuclear ribonucleoprotein particle (snRNP) binding to the intron branch site region during spliceosome assembly. We present crystal structures of S. cerevisiae Prp5p alone and in complex with ADP at 2.12 Å and 1.95 Å resolution. The three-dimensional packing of Prp5p subdomains differs strikingly from that so far observed in other DEAD-box proteins: two RecA-like subdomains adopt an "open state" conformation stabilized by extensive interactions involving sequences that flank the two subdomains. This conformation is distinct from that required for ATP hydrolysis. Consistent with this, Prp5p mutations that destabilize interdomain interactions exhibited increased ATPase activity in vitro and inhibited splicing of suboptimal branch site substrates in vivo, whereas restoration of interdomain interactions reversed these effects. We conclude that the Prp5p open state conformation is biologically relevant and that disruption of the interdomain interaction facilitates a large-scale conformational change of Prp5p during U2 snRNP-branch site recognition.
- State Key Laboratory of Bio-organic and Natural Products Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai 200032, China.
Organizational Affiliation: