Dual inhibition of HIV-1 replication by integrase-LEDGF allosteric inhibitors is predominant at the post-integration stage.
Le Rouzic, E., Bonnard, D., Chasset, S., Bruneau, J.M., Chevreuil, F., Le Strat, F., Nguyen, J., Beauvoir, R., Amadori, C., Brias, J., Vomscheid, S., Eiler, S., Levy, N., Delelis, O., Deprez, E., Saib, A., Zamborlini, A., Emiliani, S., Ruff, M., Ledoussal, B., Moreau, F., Benarous, R.(2013) Retrovirology 10: 144-144
- PubMed: 24261564 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1742-4690-10-144
- Primary Citation Related Structures:
4LH4, 4LH5 - PubMed Abstract:
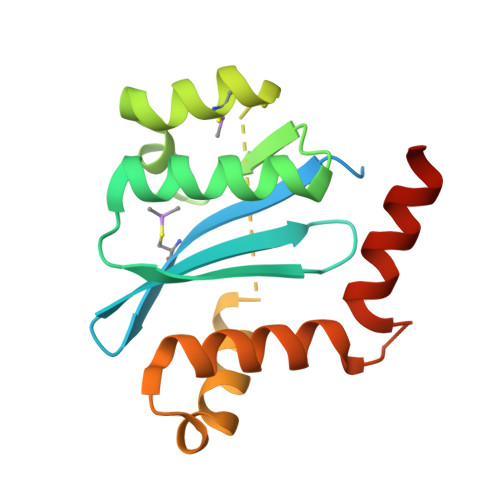
LEDGF/p75 (LEDGF) is the main cellular cofactor of HIV-1 integrase (IN). It acts as a tethering factor for IN, and targets the integration of HIV in actively transcribed gene regions of chromatin. A recently developed class of IN allosteric inhibitors can inhibit the LEDGF-IN interaction. We describe a new series of IN-LEDGF allosteric inhibitors, the most active of which is Mut101. We determined the crystal structure of Mut101 in complex with IN and showed that the compound binds to the LEDGF-binding pocket, promoting conformational changes of IN which explain at the atomic level the allosteric effect of the IN/LEDGF interaction inhibitor on IN functions. In vitro, Mut101 inhibited both IN-LEDGF interaction and IN strand transfer activity while enhancing IN-IN interaction. Time of addition experiments indicated that Mut101 behaved as an integration inhibitor. Mut101 was fully active on HIV-1 mutants resistant to INSTIs and other classes of anti-HIV drugs, indicative that this compound has a new mode of action. However, we found that Mut101 also displayed a more potent antiretroviral activity at a post-integration step. Infectivity of viral particles produced in presence of Mut101 was severely decreased. This latter effect also required the binding of the compound to the LEDGF-binding pocket. Mut101 has dual anti-HIV-1 activity, at integration and post-integration steps of the viral replication cycle, by binding to a unique target on IN (the LEDGF-binding pocket). The post-integration block of HIV-1 replication in virus-producer cells is the mechanism by which Mut101 is most active as an antiretroviral. To explain this difference between Mut101 antiretroviral activity at integration and post-integration stages, we propose the following model: LEDGF is a nuclear, chromatin-bound protein that is absent in the cytoplasm. Therefore, LEDGF can outcompete compound binding to IN in the nucleus of target cells lowering its antiretroviral activity at integration, but not in the cytoplasm where post-integration production of infectious viral particles takes place.
- Biodim Mutabilis, Romainville 93230, France. richard.benarous@mutabilis.fr.
Organizational Affiliation: