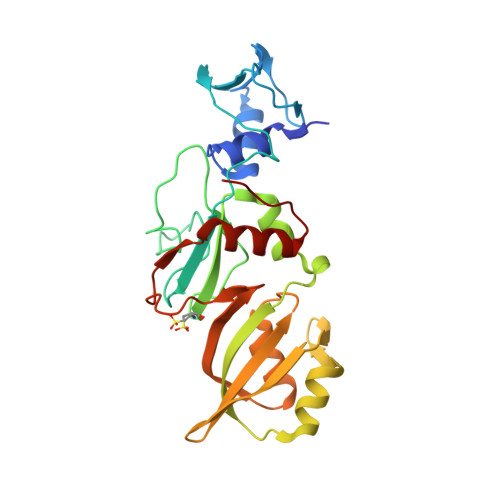
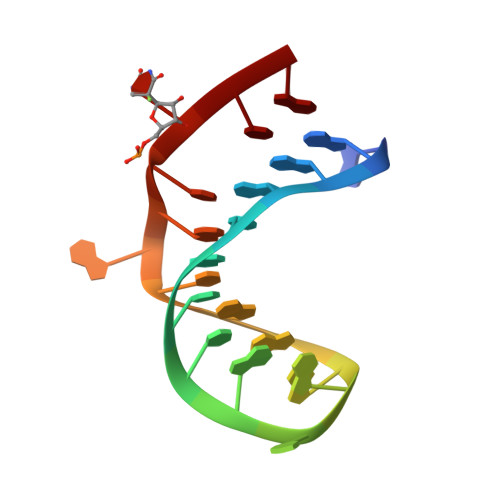
The mechanism of pseudouridine synthases from a covalent complex with RNA, and alternate specificity for U2605 versus U2604 between close homologs.
Czudnochowski, N., Ashley, G.W., Santi, D.V., Alian, A., Finer-Moore, J., Stroud, R.M.(2014) Nucleic Acids Res 42: 2037-2048
- PubMed: 24214967
- DOI: https://doi.org/10.1093/nar/gkt1050
- Primary Citation Related Structures:
4LAB, 4LGT - PubMed Abstract:
RluB catalyses the modification of U2605 to pseudouridine (Ψ) in a stem-loop at the peptidyl transferase center of Escherichia coli 23S rRNA. The homolog RluF is specific to the adjacent nucleotide in the stem, U2604. The 1.3 Å resolution crystal structure of the complex between the catalytic domain of RluB and the isolated substrate stem-loop, in which the target uridine is substituted by 5-fluorouridine (5-FU), reveals a covalent bond between the isomerized target base and tyrosine 140. The structure is compared with the catalytic domain alone determined at 2.5 Å resolution. The RluB-bound stem-loop has essentially the same secondary structure as in the ribosome, with a bulge at A2602, but with 5-FU2605 flipped into the active site. We showed earlier that RluF induced a frame-shift of the RNA, moving A2602 into the stem and translating its target, U2604, into the active site. A hydrogen-bonding network stabilizes the bulge in the RluB-RNA but is not conserved in RluF and so RluF cannot stabilize the bulge. On the basis of the covalent bond between enzyme and isomerized 5-FU we propose a Michael addition mechanism for pseudouridine formation that is consistent with all experimental data.
- Department of Biochemistry and Biophysics, University of California, San Francisco, 600 16th Street, San Francisco, CA 94158, USA, ProLynx, 455 Mission Bay Blvd., Suite 145, San Francisco, CA 94158, USA and Faculty of Biology, Technion-Israel Institute of Technology, Technion City, Haifa 320003, Israel.
Organizational Affiliation: