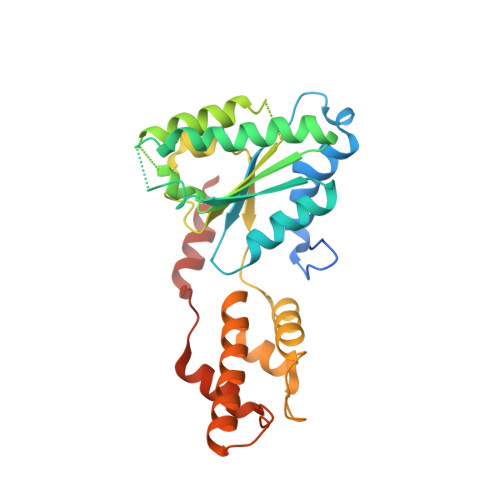
The Oligomeric State of the Active Vps4 AAA ATPase.
Monroe, N., Han, H., Gonciarz, M.D., Eckert, D.M., Karren, M.A., Whitby, F.G., Sundquist, W.I., Hill, C.P.(2014) J Mol Biology 426: 510-525
- PubMed: 24161953
- DOI: https://doi.org/10.1016/j.jmb.2013.09.043
- Primary Citation of Related Structures:
4LCB, 4LGM - PubMed Abstract:
The cellular ESCRT (endosomal sorting complexes required for transport) pathway drives membrane constriction toward the cytosol and effects membrane fission during cytokinesis, endosomal sorting, and the release of many enveloped viruses, including the human immunodeficiency virus. A component of this pathway, the AAA ATPase Vps4, provides energy for pathway progression. Although it is established that Vps4 functions as an oligomer, subunit stoichiometry and other fundamental features of the functional enzyme are unclear. Here, we report that although some mutant Vps4 proteins form dodecameric assemblies, active wild-type Saccharomyces cerevisiae and Sulfolobus solfataricus Vps4 enzymes can form hexamers in the presence of ATP and ADP, as assayed by size-exclusion chromatography and equilibrium analytical ultracentrifugation. The Vta1p activator binds hexameric yeast Vps4p without changing the oligomeric state of Vps4p, implying that the active Vta1p-Vps4p complex also contains a single hexameric ring. Additionally, we report crystal structures of two different archaeal Vps4 homologs, whose structures and lattice interactions suggest a conserved mode of oligomerization. Disruption of the proposed hexamerization interface by mutagenesis abolished the ATPase activity of archaeal Vps4 proteins and blocked Vps4p function in S. cerevisiae. These data challenge the prevailing model that active Vps4 is a double-ring dodecamer, and argue that, like other type I AAA ATPases, Vps4 functions as a single ring with six subunits.
- Department of Biochemistry, University of Utah School of Medicine, 15 North Medical Drive East RM 4100, Salt Lake City, UT 84112-5650, USA.
Organizational Affiliation: